Professional Documents
Culture Documents
SB Exercise1
Uploaded by
Imran Al NoorOriginal Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
SB Exercise1
Uploaded by
Imran Al NoorCopyright:
Available Formats
High-Level Computer Vision
Summer Semester 2013 Prof. Bernt Schiele, Dr. Mario Fritz <{schiele,mfritz}@mpi-inf.mpg.de> Dr. Fabio Galasso, Siyu Tang <{galasso,tang}@mpi-inf-mpg.de>
Exercise 1: Introduction to Matlab and Image Filtering
In this exercise you will rst familiarise yourself with the essentials of the Matlab programming and then implement several basic image ltering routines which will later be used for more complex image processing tasks such as edge detection. Download the le exercise1.tar.gz from the course web page and unpack it in a separate directory. This le contains a placeholder for each function you will have to implement in this exercise, and your task is to ll the missing code corresponding to each subproblem. The le also includes two images: graf.png and gantrycrane.png, which we will use for testing purposes. The script ex1.m sequentially executes the code corresponding to each subproblem and visualizes the results. Ideally, after you have implemented all the missing code you should be able to execute it without errors.
Question 1: Matlab Tutorial
a) Create your working directory. Run Matlab by clicking on the Matlab icon or by typing matlab in the command shell. Change the directory to your working directory. b) Watch the video Getting Started with Matlab available at http://www.mathworks.de/videos/matlab/ getting-started-with-matlab.html. c) Read the introductory parts of the Matlab guide available at www.mathworks.com/help/pdf_doc/matlab/ getstart.pdf. The chapters you should read are Matrices, Graphics and Flow Control. d) Walk through the following Matlab demos (www.mathworks.com/products/matlab/demos.html): Mathematics Basic Matrix Operations Matrix Manipulation Graphics 2-D Plots 3-D Plots Images and Matrices Programming Manipulating Multidimensional Arrays Function Functions
Question 2: Basic Image Processing
Download the le exercise1.tar.gz from the class web page and uncompress it in your working directory. This le contains two example images, which you will manipulate in the following. For the rst exercises, we provide example code. Try it out and shortly explain what happens. For the later exercises, we only provide a code framework which you should complete yourself. a) Use Matlab documentation to learn about the following commands which we will use in this exercise. imread, image, imshow, imagesc, colormap, imrotate. To obtain information about a particular command you can type help followed by the command name in the Matlab command prompt. b) Read the image graf.png and display it in a window with the command imshow. graf=imread(graf.png); imshow(graf); Convert the color image to gray values and display it with dierent Matlab commands.
graf_gray=rgb2gray(graf) close all; image(graf_gray); imagesc(graf_gray); imshow(graf_gray); c) Look at the colormap. Create and set a new colormap which varies from 1 to 0, 0 to 1, and several random maps. Display the image. Explain what happens. map=colormap; map1=map(end:-1:1,:); map2=rand(64,3); close all; image(graf_gray); colormap(map1); colormap(map2); colormap(default); colormap(gray); d) Brighten and darken the image. bgraf=graf*2; image(bgraf); dgraf=graf/2; image(dgraf); e) Assign row 200 in the image to a variable. Display it with a plot. Repeat that for column 300. row200=graf_gray(200,:); plot(row200); column300=graf_gray(:,300); plot(column300); f) Remove a rectangular region from the image and display the region. Crop a rectangular region from the image containing only the head of the karate man and display it as a new image. close all; graf1=graf; graf1(130:260,240:450, :)=0; imshow(graf1); graf2=graf(130:260,240:450,:); imshow(graf2); g) Crop the head of the chicken and rotate it by 45 degrees. Try dierent methods (nearest, bilinear, bicubic). Explain the observations. close all; subplot(1, 3, 1); graf4=imrotate(graf2,45,nearest); imshow(graf4); subplot(1, 3, 2); graf4=imrotate(graf2,45,bilinear); imshow(graf4); subplot(1, 3, 3); graf4=imrotate(graf2,45,bicubic); imshow(graf4); h) Sample the image (reduce the size). Use dierent sampling step sizes (2, 4, 8). Explain what happens. close all; subplot(1, 3, 1); graf5=graf_gray(1:2:end,1:2:end); imshow(graf5); subplot(1, 3, 2); graf5=graf_gray(1:4:end,1:4:end); imshow(graf5); subplot(1, 3, 3); graf5=graf_gray(1:8:end,1:8:end); imshow(graf5); i) Create a mirror image. Concatenate 2 or 4 images. graf6=graf_gray(:,end:-1:1); imshow(graf6);
graf7=[graf_gray,graf_gray;graf_gray,graf_gray]; imshow(graf7); j) Assign row 200 in the image to a variable. Display it with a plot. Create a lter which is a vector of 10 equal elements summing to 1. Convolve the row with the lter and plot the result. Explain what happens. row200=graf_gray(200,:); plot(row200); filt1=ones(1,10)/10; newrow=conv(double(row200), filt1); hold on; plot(newrow, r-); k) Create a 2D blur lter of size 10 10 which sums to 1. Convolve it with the image and display the new image. Create a 1D lter of size 10 which sums to 1. Convolve it with the lines of the image. Transpose the lter and convolve it with rows of the ltered image. Explain what happens. filtersize=10; filt2D=ones(filtersize,filtersize)/(filtersize*filtersize); graf8=conv2(double(graf_gray),filt2D); imagesc(graf8); filt1D=ones(1,filtersize)/filtersize; graf9=conv2(double(graf_gray),filt1D); graf10=conv2(graf9,filt1D); imagesc(graf10); max(max(graf10 - graf8)) l) Create a function called basicimproc which takes as arguments an image inimg and ve numbers x, y, the size of a square, rotationangle and filtersize. The function should crop a square region out of the image, rotate it by the desired angle, blur the cropped region, and then return it as outimg. It should also display the intermediate result images. Create a new le called basicimproc.m with your preferred editor and begin the script: function outimg=basicimproc(inimg, x, y, size, rotationangle, filtersize) ... ... end
Question 3: Image Filtering (15 points)
a) Implement a method which computes the values of a Gaussian for a given variance and a number of samples. G= x2 1 exp( 2 ) 2 2 (1)
where is the standard deviation and x is a vector of integer values. Open the le gauss.m with your preferred editor and begin the script: function G=gauss(sigma) ... ... end b) Implement a 2D Gaussian lter in gaussianfilter.m. The function should take an image as an input and return the result of convolution of this image with 2D Gaussian kernel. See Fig. 1 for illustration of Gaussian ltering. Width and height of the lter should be 2 (3 sigma) + 1. You can take advantage of the Matlabs conv2 function if you dont want to implement convolution yourself. Open the le called gaussianfilter.m with your preferred editor and begin the script: function outimg=gaussianfilter(inimg,sigma) ... ... end
(a)
(b)
Figure 1: (a) Original image (b) Image after applying a Gaussian lter with = 4.0. Hint: use the fact that 2D Gaussian lter is separable to speed up computations. c) Implement a function gaussdx.m for creating a Gaussian derivative lter in 1D according to the following equation d G = dx = x2 d 1 exp( 2 ) dx 2 2 1 x2 x exp( 2 ) 2 2 3
(2) (3)
The eect of applying a lter can be studied by observing its so-called impulse response. For this, create a test image in which only the central pixel has a non-zero value: imgImp = zeros(25,25); imgImp(13,13) = 1.0; Now, create the following 1D lter kernels G and D. sigma = 6.0; G = gauss(sigma); D = gaussdx(sigma); What happens when you apply the following lter combinations? (For best visualization, display the result image with the imagesc command). 1. 2. 3. 4. 5. rst rst rst rst rst G, then G. G, then D. D, then G. G, then D. D, then G.
d) Use the functions gauss and gaussdx in order to implement a function gaussderiv that returns the 2D Gaussian derivatives of an input image in x and y direction. Try the function on the given example images. Please turn in your solution by sending an email to Fabio Galasso <galasso@ mpi-inf. mpg. de> including all relevant m-les before Wednesday, May 1st, 23:59.
You might also like
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryFrom EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryRating: 3.5 out of 5 stars3.5/5 (231)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)From EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Rating: 4.5 out of 5 stars4.5/5 (119)
- Never Split the Difference: Negotiating As If Your Life Depended On ItFrom EverandNever Split the Difference: Negotiating As If Your Life Depended On ItRating: 4.5 out of 5 stars4.5/5 (838)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaFrom EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaRating: 4.5 out of 5 stars4.5/5 (265)
- The Little Book of Hygge: Danish Secrets to Happy LivingFrom EverandThe Little Book of Hygge: Danish Secrets to Happy LivingRating: 3.5 out of 5 stars3.5/5 (399)
- Grit: The Power of Passion and PerseveranceFrom EverandGrit: The Power of Passion and PerseveranceRating: 4 out of 5 stars4/5 (587)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyFrom EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyRating: 3.5 out of 5 stars3.5/5 (2219)
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeFrom EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeRating: 4 out of 5 stars4/5 (5794)
- Team of Rivals: The Political Genius of Abraham LincolnFrom EverandTeam of Rivals: The Political Genius of Abraham LincolnRating: 4.5 out of 5 stars4.5/5 (234)
- Shoe Dog: A Memoir by the Creator of NikeFrom EverandShoe Dog: A Memoir by the Creator of NikeRating: 4.5 out of 5 stars4.5/5 (537)
- The Emperor of All Maladies: A Biography of CancerFrom EverandThe Emperor of All Maladies: A Biography of CancerRating: 4.5 out of 5 stars4.5/5 (271)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreFrom EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreRating: 4 out of 5 stars4/5 (1090)
- Her Body and Other Parties: StoriesFrom EverandHer Body and Other Parties: StoriesRating: 4 out of 5 stars4/5 (821)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersFrom EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersRating: 4.5 out of 5 stars4.5/5 (344)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceFrom EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceRating: 4 out of 5 stars4/5 (890)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureFrom EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureRating: 4.5 out of 5 stars4.5/5 (474)
- The Unwinding: An Inner History of the New AmericaFrom EverandThe Unwinding: An Inner History of the New AmericaRating: 4 out of 5 stars4/5 (45)
- The Yellow House: A Memoir (2019 National Book Award Winner)From EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Rating: 4 out of 5 stars4/5 (98)
- On Fire: The (Burning) Case for a Green New DealFrom EverandOn Fire: The (Burning) Case for a Green New DealRating: 4 out of 5 stars4/5 (73)
- SCM NotesDocument29 pagesSCM NotesNisha Pradeepa100% (2)
- TG Comply With WP Hygiene Proc 270812 PDFDocument224 pagesTG Comply With WP Hygiene Proc 270812 PDFEmelita MendezNo ratings yet
- Unit 8 Risk in The WorkplaceDocument11 pagesUnit 8 Risk in The WorkplaceAnonymous WalvB8No ratings yet
- RA 8042 and RA 10022 ComparedDocument37 pagesRA 8042 and RA 10022 ComparedCj GarciaNo ratings yet
- 15 04 06 SCDocument30 pages15 04 06 SCSugarNo ratings yet
- MODEL QUESTION PAPER OF HRM Open CourceDocument2 pagesMODEL QUESTION PAPER OF HRM Open CourceTitus Clement100% (3)
- Q.850 ISDN Cause CodesDocument12 pagesQ.850 ISDN Cause CodesJack CastineNo ratings yet
- Unit 06 Extra Grammar ExercisesDocument3 pagesUnit 06 Extra Grammar ExercisesLeo Muñoz43% (7)
- Energy Efficiency Existing Ship Index (Eexi) : Regulatory DebriefDocument8 pagesEnergy Efficiency Existing Ship Index (Eexi) : Regulatory DebriefSalomonlcNo ratings yet
- AcousticsDocument72 pagesAcousticsImran Al NoorNo ratings yet
- UCLA - Above The Clouds A Berkeley View of Cloud Computing (08-2012)Document25 pagesUCLA - Above The Clouds A Berkeley View of Cloud Computing (08-2012)Angela Hernández HerreraNo ratings yet
- PESQ - A New Method for Speech Quality AssessmentDocument4 pagesPESQ - A New Method for Speech Quality AssessmentImran Al NoorNo ratings yet
- Multi-Fidelity UI SpecificationsDocument16 pagesMulti-Fidelity UI SpecificationsImran Al NoorNo ratings yet
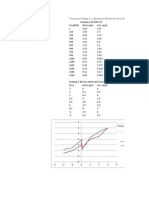
- Frequency/Voltage vs. Capacitance Measurement in OLED's (Stressed and Distressed)Document3 pagesFrequency/Voltage vs. Capacitance Measurement in OLED's (Stressed and Distressed)Imran Al NoorNo ratings yet
- E-Speaking Voice and Speech Recognition CommandsDocument3 pagesE-Speaking Voice and Speech Recognition CommandsZuhairi SyazwanNo ratings yet
- Things To ConsiderDocument1 pageThings To ConsiderImran Al NoorNo ratings yet
- Free-hand 3D modeling interface reduces cognitive loadDocument11 pagesFree-hand 3D modeling interface reduces cognitive loadImran Al NoorNo ratings yet
- Leyendo Matemáticas - ArticuloDocument3 pagesLeyendo Matemáticas - ArticuloJulio Cesar SaNo ratings yet
- Automation Systems Discrete Event ControlDocument13 pagesAutomation Systems Discrete Event ControlImran Al NoorNo ratings yet
- 5.doppler Diversity For OFDM Wireless Mobile Communications. Part IIDocument4 pages5.doppler Diversity For OFDM Wireless Mobile Communications. Part IIImran Al NoorNo ratings yet
- Lesson 2: Mathematics Review: Spring 2006 Process Dynamics, Operations, and Control 10.450Document11 pagesLesson 2: Mathematics Review: Spring 2006 Process Dynamics, Operations, and Control 10.450Imran Al NoorNo ratings yet
- 1 s2.0 S0313592622001369 MainDocument14 pages1 s2.0 S0313592622001369 MainNGOC VO LE THANHNo ratings yet
- Marking SchemeDocument8 pagesMarking Schememohamed sajithNo ratings yet
- Holmes 1993Document8 pagesHolmes 1993Rumaisa KrubaNo ratings yet
- CANDIDATE'S BIO DATADocument2 pagesCANDIDATE'S BIO DATAAamir ArainNo ratings yet
- Research Paper About Cebu PacificDocument8 pagesResearch Paper About Cebu Pacificwqbdxbvkg100% (1)
- What is Software Development Life Cycle (SDLC)? Key Phases and ActivitiesDocument11 pagesWhat is Software Development Life Cycle (SDLC)? Key Phases and ActivitiessachinNo ratings yet
- Microsoft Windows 98 Second Edition README For Tips and Tricks, April 1999Document8 pagesMicrosoft Windows 98 Second Edition README For Tips and Tricks, April 1999scriNo ratings yet
- Elegant Tranquil Blue Agency by SlidesgoDocument41 pagesElegant Tranquil Blue Agency by SlidesgoJoana TavaresNo ratings yet
- Case Study ON: The Spark Batteries LTDDocument8 pagesCase Study ON: The Spark Batteries LTDRitam chaturvediNo ratings yet
- Instructions Manual Skatey 150/250/400/600Document19 pagesInstructions Manual Skatey 150/250/400/600Denys GavrylovNo ratings yet
- Section 2 in The Forest (Conservation) Act, 1980Document1 pageSection 2 in The Forest (Conservation) Act, 1980amit singhNo ratings yet
- SOLUTIONS : Midterm Exam For Simulation (CAP 4800)Document14 pagesSOLUTIONS : Midterm Exam For Simulation (CAP 4800)Amit DostNo ratings yet
- Rheomix 141Document5 pagesRheomix 141Haresh BhavnaniNo ratings yet
- Essential earthquake preparedness stepsDocument6 pagesEssential earthquake preparedness stepsRalphNacisNo ratings yet
- Lecture 2 Leader-Centred PerspectivesDocument24 pagesLecture 2 Leader-Centred PerspectivesLIVINGSTONE CAESARNo ratings yet
- Nikita Project 01-06-2016Document38 pagesNikita Project 01-06-2016Shobhit GoswamiNo ratings yet
- Project CST 383Document1,083 pagesProject CST 383api-668525404No ratings yet
- ION8650 DatasheetDocument11 pagesION8650 DatasheetAlthaf Axel HiroshiNo ratings yet
- Norlys 2016Document124 pagesNorlys 2016elektrospecNo ratings yet
- Loctite 270™: Technical Data SheetDocument4 pagesLoctite 270™: Technical Data SheetM Jobayer AzadNo ratings yet
- Guidelines Regarding The Handling of Cable Drums During Transport and StorageDocument5 pagesGuidelines Regarding The Handling of Cable Drums During Transport and StorageJegan SureshNo ratings yet