Professional Documents
Culture Documents
PNAS 2003 Al Hajj 3983 8
Uploaded by
mememe123123Original Title
Copyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
PNAS 2003 Al Hajj 3983 8
Uploaded by
mememe123123Copyright:
Available Formats
Corrections
NEUROSCIENCE. For the article Histone deacetylase inhibitors
prevent oxidative neuronal death independent of expanded
polyglutamine repeats via an Sp1-dependent pathway, by Hoon
Ryu, Junghee Lee, Beatrix A. Olofsson, Aziza Mwidau, Alpaslan
Deodoglu, Maria Escudero, Erik Flemington, Jane AzizkhanClifford, Robert J. Ferrante, and Rajiv R. Ratan, which appeared in issue 7, April 1, 2003, of Proc. Natl. Acad. Sci. USA
(100, 42814286; First Published March 14, 2003; 10.1073
pnas.0737363100), the author name Alpaslan Deodoglu should
have appeared as Alpaslan Dedeoglu. The corrected author line
appears below. The online version has been corrected.
Hoon Ryu, Junghee Lee, Beatrix A. Olofsson,
Aziza Mwidau, Alpaslan Dedeoglu, Maria Escudero,
Erik Flemington, Jane Azizkhan-Clifford,
Robert J. Ferrante, and Rajiv R. Ratan
www.pnas.orgcgidoi10.1073pnas.1231735100
COMMENTARY. For the article What happens to genes in duplicated genomes, by Elizabeth A. Kellogg, which appeared
in issue 8, April 15, 2003, of Proc. Natl. Acad. Sci. USA
(100, 4369 4371; First Published April 7, 2003; 10.1073
pnas.0831050100), the pull quote in the second column on page
4370 read Subfunctionalization occurs in some genes and is not
an immediate product of polyploidization. It should have read
Subfunctionalization occurs in some genes and is an immediate
product of polyploidization. This error occurred during the
editorial process and is not the fault of the author. PNAS regrets
this error.
CELL BIOLOGY. For the article Prospective identification of tumorigenic breast cancer cells, by Muhammad Al-Hajj, Max S.
Wicha, Adalberto Benito-Hernandez, Sean J. Morrison, and
Michael F. Clarke, which appeared in issue 7, April 1, 2003, of
Proc. Natl. Acad. Sci. USA (100, 39833988; First Published
March 10, 2003; 10.1073pnas.0530291100), the authors note
that the following statement was inadvertently omitted from the
acknowledgements: The results of this study support a patentpending technology that is exclusively licensed to Cancer Stem
Cell Genomics (CSCG) in which the authors and the University
of Michigan have a financial interest.
www.pnas.orgcgidoi10.1073pnas.1131491100
IMMUNOLOGY. For the article Antigen presentation by keratin-
ocytes directs autoimmune skin disease, by Lian Fan, Brian W.
Busser, Traci Q. Lifsted, David Lo, and Terri M. Laufer, which
appeared in issue 6, March 18, 2003, of Proc. Natl. Acad. Sci. USA
(100, 33863391; First Published March 10, 2003; 10.1073
pnas.0437899100), the authors request that Mohamed Oukka,
Department of Immunology and Infectious Diseases, Harvard
School of Public Health, Boston, MA 02115, be added to the list
of authors between Traci Q. Lifsted and David Lo. The revised
author line appears below.
Lian Fan, Brian W. Busser, Traci Q. Lifsted,
Mohamed Oukka, David Lo, and Terri M. Laufer
www.pnas.orgcgidoi10.1073pnas.1232295100
www.pnas.orgcgidoi10.1073pnas.1232296100
6890 6891 PNAS May 27, 2003 vol. 100 no. 11
www.pnas.org
10.1073/pnas.0831001100), in Fig. 1 A the hydroxyl group in
position 17 for 5-androstenediol (Adiol), testosterone, and
dihydrotestosterone (DHT) should be connected with a single,
rather than a double, bond. The conclusions are unchanged by
this typographical error. The corrected figure and its legend
appear below.
CORRECTIONS
BIOCHEMISTRY. For the article 3-Acetoxyandrost-1,5-diene17-ethylene ketal functions as a potent antiandrogen with
marginal agonist activity, by Hiroshi Miyamoto, Padma
Marwah, Ashok Marwah, Henry Lardy, and Chawnshang
Chang, which appeared in issue 8, April 15, 2003, of Proc. Natl.
Acad. Sci. USA (100, 4440 4444; First Published April 2, 2003,
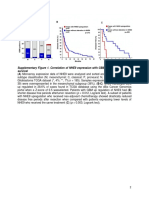
Fig. 1. The structures of DHEA derivatives and effects on AR transcriptional activity. (A) The structures of compounds nos. 5, 10, 14, 15, 16, and 17, DHEA, Adiol,
testosterone, and DHT. (B) PC-3 cells were transfected with the WT AR expression plasmid pSG5-AR and MMTV-Luc. After transfection, cells were cultured for
24 h with 1 nM DHT or 1,000 nM of various DHEA derivatives. The Luc activity is presented relative to that of EtOH treatment (white bar; set as 1-fold). Values
represent the mean SD of at least three determinations. (C) PC-3 cells were transfected with the pSG5-AR and MMTV-Luc. After transfection, cells were cultured
for 24 h with various concentrations of compounds nos. 5, 10 (ADEK), 14, or 16 in the presence of 1 nM DHT. The Luc activity is presented relative to that in the
presence of DHT (black bar; set as 100%). Values represent the mean SD of at least three determinations.
www.pnas.orgcgidoi10.1073pnas.1232068100
PNAS May 27, 2003 vol. 100 no. 11 6891
SEE COMMENTARY
Prospective identification of tumorigenic breast
cancer cells
Muhammad Al-Hajj*, Max S. Wicha*, Adalberto Benito-Hernandez, Sean J. Morrison*, and Michael F. Clarke*
Departments of *Internal Medicine and Pathology, Comprehensive Cancer Center, Department of Developmental Biology, and Howard Hughes Medical
Institute, University of Michigan Medical School, Ann Arbor, MI 48109
Communicated by Jack E. Dixon, University of Michigan Medical School, Ann Arbor, MI, January 16, 2003 (received for review December 18, 2002)
espite advances in detection and treatment of metastatic
breast cancer, mortality from this disease remains high
because current therapies are limited by the emergence of
therapy-resistant cancer cells (1, 2). As a result, metastatic breast
cancer remains an incurable disease by current treatment strategies. Cancers are believed to arise from a series of sequential
mutations that occur as a result of genetic instability andor
environmental factors (3, 4). A better understanding of the
consequences of these mutations on the underlying biology of
the neoplastic cells may lead to new therapeutic strategies.
In solid tumors, it has been demonstrated that only a small
proportion of the tumor cells are able to form colonies in an in vitro
clonogenic assay (511). Furthermore, large numbers of cells must
typically be transplanted to form tumors in xenograft models. One
possible explanation for these observations is that every cell within
a tumor has the ability to proliferate and form new tumors but that
the probability of an individual cell completing the necessary steps
in these assays is small. An alternative explanation is that only a rare,
phenotypically distinct subset of cells has the capacity to significantly proliferate and form new tumors, but that cells within this
subset do so very efficiently (12). To distinguish between these
possibilities, it is necessary to identify the clonogenic cells in these
tumors with markers that distinguish these cells from other nontumorigenic cells. This identification has been accomplished in
acute myelogenous leukemia, where it was demonstrated that a
specific subpopulation of leukemia cells (that expressed markers
similar to normal hematopoietic stem cells) was consistently enriched for clonogenic activity in nonobese diabeticsevere combined immunodeficient (NODSCID) immunocompromised mice,
whereas other cancer cells were depleted of clonogenic activity
(1315). Such experiments have not been reported in solid cancers.
If this model were also true for solid tumors, and only a small subset
of cells within a tumor possess the capacity to proliferate and form
new tumors, this finding would have significant implications for
www.pnas.orgcgidoi10.1073pnas.0530291100
understanding the biology of and developing therapeutic strategies
for these neoplasms.
To investigate the mechanisms of solid tumor heterogeneity, we
developed a modification of the NODSCID mouse model in which
human breast cancers were efficiently propagated in the mouse
mammary fat pad (16). In the present study, we show that solid
tumors contain a distinct population of cells with the exclusive
ability to form tumors in mice. We refer to these cells as tumorigenic
cells, or cancer-initiating cells, because they consistently formed
tumors, whereas other cancer cell populations were depleted of cells
capable of tumor formation. We identified cell surface markers that
can distinguish between these cell populations. Our findings provide a previously uncharacterized model of breast tumor biology in
which a defined subset of cells drives tumorigenesis, as well as
generating tumor cell heterogeneity. The prospective identification
of this tumorigenic population of cancer cells should allow for the
identification of molecules expressed in these cells that could serve
as targets to eliminate this critical population of cancer cells.
Materials and Methods
Mouse Preparation. Eight-week-old female NODSCID mice
were anesthetized by an i.p. injection of 0.2 ml of ketamine
xylazine (300 mg ketamine combined with 20 mg of xylazine in
a 4-ml volume; 0.02 ml of the solution was used per 20-g mouse).
Dilution to 200 l was done by using Hanks balanced salt
solution (HBSS). Mice were then treated with VP-16 (etoposide)
via an i.p. injection (30-mg etoposide dose per 1-kg mouse,
diluted in serum-free HBSS for a final injection volume of 200
l). At the same time, estrogen pellets were placed s.c. on the
back of the mouses neck by using a trocar. All tumor injections
implants were done 5 days after this procedure. In the following
procedures, mice were always anesthetized as described above.
Primary Tumor Specimen Implantations. For the implantation of
fresh specimens, samples of human breast tumors were received
within an hour after surgery. The tumors were cut up with scissors
into small pieces, and the pieces were then minced with a blade to
yield 2 2-mm pieces. Mincing was done in sterile RPMI medium
1640 under sterile conditions on ice. The tumor pieces were washed
with serum-free HBSS before implantation. A 2-mm incision was
then made in the mid abdomen area, and by using a trocar, one to
two small tumor pieces were implanted in the region of the upper
right and upper left mammary fat pads (right below the second
nipple on both sides). A 6-0 suture was wrapped twice around the
mammary fat pad nipple, allowing it to hold the implanted pieces
in place. Sutures were removed after 5 days. Nexaban was used to
seal the incision, and mice were monitored weekly for tumor
growth.
Pleural Effusion Injections. For the injection of the pleural effu-
sions, cells were received shortly after thoracentesis and washed
Abbreviations: NODSCID, nonobese diabeticsevere combined immunodeficient; HICS,
heat-inactivated calf serum; ESA, epithelial-specific antigen; Tn, tumor n.
See commentary on page 3547.
To
whom correspondence should be addressed. E-mail: mclarke@med.umich.edu.
PNAS April 1, 2003 vol. 100 no. 7 39833988
CELL BIOLOGY
Breast cancer is the most common malignancy in United States
women, accounting for >40,000 deaths each year. These breast
tumors are comprised of phenotypically diverse populations of breast
cancer cells. Using a model in which human breast cancer cells were
grown in immunocompromised mice, we found that only a minority
of breast cancer cells had the ability to form new tumors. We were
able to distinguish the tumorigenic (tumor initiating) from the nontumorigenic cancer cells based on cell surface marker expression. We
prospectively identified and isolated the tumorigenic cells as
CD44CD24/lowLineage in eight of nine patients. As few as 100 cells
with this phenotype were able to form tumors in mice, whereas tens
of thousands of cells with alternate phenotypes failed to form
tumors. The tumorigenic subpopulation could be serially passaged:
each time cells within this population generated new tumors containing additional CD44CD24/lowLineage tumorigenic cells as well
as the phenotypically diverse mixed populations of nontumorigenic
cells present in the initial tumor. The ability to prospectively identify
tumorigenic cancer cells will facilitate the elucidation of pathways
that regulate their growth and survival. Furthermore, because these
cells drive tumor development, strategies designed to target this
population may lead to more effective therapies.
Table 1. Engraftment of human breast cancers into NODSCID mice
Tumor
T1
T2
T3
T4
T5
T6
T7
T8
T9
Origin
Formation in
mice
Passage in
mice
Diagnosis
Metastasis
Breast primary
Metastasis
Metastasis
Metastasis
Metastasis
Metastasis
Metastasis
Metastasis
Yes
Yes
Yes
Yes
Yes
Yes
Yes
Yes
Yes
Yes
Yes
Yes
No
Yes
Yes
Yes
Yes
Yes
Infiltrating ductal carcinoma
Adenocarcinoma
Invasive lobular carcinoma
Invasive lobular carcinoma
Invasive lobular carcinoma
Inflammatory breast carcinoma
Invasive lobular carcinoma
Inflammatory breast carcinoma
Adenocarcinoma
Mice were injected with unsorted T1 and T3 cells and a 2-mm piece of T2. Cells from T4 T9 were isolated by
flow cytometry as described in Fig. 1. All nine tumors tested engrafted in our NODSCID mouse model. Except for
T2, which was a primary breast tumor, all other tumors were metastases. All of the tumors were passaged serially
in mice except for T4.
with serum-free HBSS. Cells were then suspended in serum
free-RPMIMatrigel mixture (1:1 volume) and then injected
into the upper right and left mammary pads by using a 22-gauge
needle. An amount equal to 0.2 ml, containing 12 million cells,
was typically injected. The site of the needle injection was sealed
with Nexaban to prevent any cell leakage.
volume) and injected into the area of the mammary fat pad as
described above. Nexaban was used to seal the injection site.
Cell Staining for Flow Cytometry. Cells were counted and then
transferred to a 5-ml tube, washed twice with HBSS with 2%
heat-inactivated calf serum (HICS; 5 min at 1,000 rpm), then
resuspended in 100 l (per 106 cells) of HBSS with 2% HICS. Five
Preparation of Single Cell Suspensions of Tumor Cells. Before diges-
tion with collagenase, xenograft tumors or primary human tumors
were cut up into small pieces and then minced completely by using
sterile blades. To obtain single cell suspensions, either pleural
effusion cells or the resulting tumor pieces were then mixed with
ultra-pure collagenase III in medium 199 (200250 units of collagenase per ml) and allowed to incubate at 37C for 34 h. Pipetting
with a 10-ml pipette was done every 1520 min. At the end of the
incubation, cells were filtered through a 45-l nylon mesh and
washed with RPMI20% FBS, then washed twice with HBSS. Cells
to be injected were then suspended in HBSSMatrigel mix (1:1
Table 2. Tumor formation ability of sorted cells
Tumorsinjections
5 105
2 105
02
22
02
22
02
22
02
22
16
66
02
22
02
22
02
22
02
22
16
66
8
Passaged T1
CD44
CD44
B38.1
B38.1
CD24
CD24
Passaged T2
CD44
CD44
B38.1
B38.1
CD24
CD24
105
Cells were isolated by flow cytometry as described in Fig. 1 based on
expression of the indicated marker and assayed for the ability to form tumors
after injection into the mammary fat pads of NODSCID mice at 8 105, 5
105, and 2 105 cells per injection. For 12 wk, mice were examined weekly for
tumors by observation and palpation; then all mice were necropsied to look
for growths at injection sites that were too small to palpate. The number of
tumors that formed and the number of injections that were performed are
indicated for each population. All tumors were readily apparent by visual
inspection and palpation except for tumors from the CD24 population,
which were detected only upon necropsy.
3984 www.pnas.orgcgidoi10.1073pnas.0530291100
Fig. 1. Isolation of tumorigenic cells. Flow cytometry was used to isolate
subpopulations of T1 (a and b), T3 (c), T5 (d), T6 (e), and T7 ( f) cells that were
tested for tumorigenicity in NODSCID mice. T1 (b) and T3 (c) had been
passaged (P) once in NODSCID mice, whereas the rest of the cells were frozen
or unfrozen samples obtained directly after removal from a patient (UP). Cells
were stained with antibodies against CD44, CD24, Lineage markers, and
mouse-H2K (for passaged tumors obtained from mice), and 7AAD. Dead cells
(7AAD), mouse cells (H2K), and Lineage normal cells were eliminated from
all analyses. Each plot depicts the CD24 and CD44 staining patterns of live
human Lineage cancer cells, and the frequency of the boxed tumorigenic
cancer population as a percentage of cancer cells in each specimen is shown.
Al-Hajj et al.
SEE COMMENTARY
Table 3. Tumorigenic breast cancer cells were highly enriched in the ESACD44CD24/low population
Tumorsinjections
Mouse passage 1
Unsorted
CD44CD24
CD44CD24/low
CD44CD24/lowESA
CD44CD24/lowESA
Mouse passage 2
CD44CD24
CD44CD24/low
Patients tumor cells
CD44CD24
CD44CD24/low
CD44CD24/lowESA
CD44CD24/lowESA
5 105
105
5 104
2 104
104
5 103
103
500
200
100
88
88
1010
010
1010
312
010
1010
014
1414
012
010
1010
1010*
010*
44
04
44
04
16
06
09
99
03
33
04
44
08
113
1113
02
11
22
22
22
02
microliters of Sandoglobin solution (1 mgml) was then added and
incubated on ice for 10 min, after which the sample was washed
twice with HBSS2% HICS and resuspended in 100 l (per 106
cells) of HBSS2% HICS. Antibodies (appropriate dilution per
antibody) were then added and incubated for 20 min on ice, and
then washed twice with HBSS2% HICS. When needed, a secondary antibody addition was conducted by resuspending in 100 l
(per 106 cells) of HBSS2% HICS, and then adding 14 l of
secondary antibody (depending on the secondary antibody and its
concentration), followed by a 20-min incubation. When a streptavidin step was used, cells were resuspended in 100 l (per 106 cells)
of HBSS2% HICS, and then 1 l of streptavidin, conjugated with
the indicated fluorescent dye, was added, followed by a 20-min
incubation. The cells were washed twice with HBSS2% HICS and
resuspended in 0.5 ml (per million cells) of HBSS2% HICS
that contained 7-aminoactinomycin D (7AAD, 1 gml final
concentration).
Flow Cytometry. The antibodies used were anti-CD44 [allophycocyanin (APC), phycoerythrin (PE), or biotin], anti-CD24 (PE or
FITC), anti-B38.1 (APC), anti-epithelial-specific antigen (ESA)
FITC (Biomeda, Foster City, CA), and anti-H2Kd, (PharMingen).
Lineage marker antibodies were anti-CD2, -CD3 -CD10, -CD16,
-CD18, -CD31, -CD64, and -CD140b. Unless noted, antibodies
were purchased from PharMingen. Antibodies were directly conjugated to various fluorochromes, depending on the experiment. In
all experiments, mouse cells andor Lineage cells were eliminated
by discarding H2Kd (mouse histocompatibility class I) cells or
Lineage cells during flow cytometry. Dead cells were eliminated
by using the viability dye 7AAD. Flow cytometry was performed on
a FACSVantage (Becton Dickinson). Side scatter and forward
scatter profiles were used to eliminate cell doublets. Cells were
routinely sorted twice, and the cells were reanalyzed for purity,
which typically was 95%.
Results and Discussion
Tumor Specimens and Engraftment Rate. Human breast cancer
specimens obtained from primary or metastatic sites in nine
Al-Hajj et al.
different patients [designated tumors 19 (T1T9)] all engrafted in
the NODSCID mice (Table 1). In one case, the cancer cells were
obtained from a primary breast tumor (T2) whereas in other cases
the cells were obtained from metastatic pleural effusions (T1 and
T3T9). Some experiments were conducted on cells after they had
been passaged once or twice in mice (designated passage 1 and 2)
whereas other experiments were conducted on unpassaged fresh or
frozen tumor samples obtained directly from patients. During use
of human cancer cells from tumors passaged in mice, contaminating
mouse cells were removed by eliminating H2K cells (mouse
histocompatibility class I).
Identification of Tumorigenicity Markers. Breast cancer cells were
heterogeneous with respect to expression of a variety of cell surface
markers (including CD44, CD24, and B38.1). CD24 and CD44 are
adhesion molecules whereas B38.1 has been described as a breast
ovarian cancer-specific marker (1719). To determine whether
these markers could distinguish tumorigenic from nontumorigenic
cells, flow cytometry was used to isolate cells that were positive or
negative for each marker from passage 1 T1 or T2 cells. When 2
105 to 8 105 cells of each population were injected, all injections
of CD44 cells (88), B38.1 cells (88), or CD24/low cells (1212)
gave rise to visible tumors within 12 wk of injection, but none of the
CD44 cell (08), or B38.1 cell (08) injections formed detectable
tumors (Table 2). Although no tumors could be detected by
palpation in the locations injected with CD24 cells, 2 of 12 mice
injected with CD24 cells did contain small growths at the injection
site that were detected on necropsy. These growths most likely arose
from the 13% of CD24 cells that invariably contaminate the
sorted CD24 cells or alternatively from CD24 cells with reduced
proliferative capacity (Table 2). Because the CD44 cells were
exclusively B38.1, we focused on the CD44 and CD24 markers in
subsequent experiments.
Several antigens associated with normal cell types (Lineage
markers: CD2, CD3, CD10, CD16, CD18, CD31, CD64, and
CD140b) were found not to be expressed by the cancer cells
based on analyses of tumors that had been passaged multiple
times in mice. By eliminating Lineage cells from unpassaged or
PNAS April 1, 2003 vol. 100 no. 7 3985
CELL BIOLOGY
Cells were isolated from passage 1 (Mouse passage 1) T1, T2, and T3, passage 2 (Mouse passage 2) T3, and unpassaged (Patients tumor cells) T1, T4, T5, T6,
T8, and T9. CD44CD24Lineage populations and CD44CD24/lowLineage cells were isolated by flow cytometry as described in Fig. 1. The indicated number
of cells of each phenotype was injected into the breast of NODSCID mice. The frequency of tumorigenic cells calculated by the modified maximum likelihood
analysis method is 5105 if single tumorigenic cells were capable of forming tumors, and every transplanted tumorigenic cell gave rise to a tumor (33). Therefore,
this calculation may underestimate the frequency of the tumorigenic cells because it does not take into account cell cell interactions and local environmental
factors that may influence engraftment. In addition to the markers that are shown, all sorted cells in all experiments were Lineage, and the tumorigenic cells
from T1, T2, and T3 were further selected as B38.1. The mice were observed weekly for 4 6.5 mo, or until the mice became sick from the tumors.
*Two thousand cells were injected in these experiments.
Tumor formation by 5,000 T5 ESACD44CD24/lowLineage cells was detected 8 9 wk after injections, whereas tumor formation by 5,000 T5
ESACD44CD24/lowLineage cells was detected 10 12 wk after injections.
Fig. 2. DNA content of tumorigenic and nontumorigenic breast cancer cells.
The cell cycle status of the ESACD44CD24/lowLineage tumorigenic cells (a)
and the remaining Lineage nontumorigenic cancer cells (b) isolated from
T1 were determined by Hoechst 33342 staining of DNA content (20). The
tumorigenic and nontumorigenic cell populations exhibited similar cell cycle
distributions.
early passage tumor cells, normal human leukocytes, endothelial
cells, mesothelial cells, and fibroblasts were eliminated. By
microscopic examination, the Lineage tumor cells consistently
had the appearance of neoplastic cells (data not shown).
Depending on the tumor, 1135% of the Lineage cancer cells
in tumors or pleural effusions were CD44CD24/low (Fig. 1).
CD44CD24/lowLineage cells or other populations of Lineage
cancer cells that had been isolated from nine patients were injected
into the mammary fat pads of mice (Table 3). When injecting
unsorted, passaged T1 or T2 cells, 5 104 cells consistently gave rise
to tumors, but 104 cells gave rise to tumors in only a minority of
cases. In contrast, as few as 103 T1 or T2 CD44CD24/lowLineage
cells gave rise to tumors in all cases (Table 3). In T1 and T2, up to
2 104 cells that were CD44Lineage but CD24 failed to form
tumors. These data suggest that the CD44CD24/lowLineage
population is 10- to 50-fold enriched for the ability to form tumors
in NODSCID mice relative to unfractionated tumor cells.
Whether the CD44CD24/lowLineage cells were isolated from
passaged tumors (T1, T2, and T3) or from unpassaged cancer cells
obtained directly from patients (T1, T4T6, T8, and T9), they were
enriched for tumorigenic activity. Note that T7 was the only one of
nine cancers studied that did not fit this pattern (Fig. 1f ). Other than
T7, CD24Lineage cancer cells in both unpassaged and passaged
tumors were unable to form new tumors (Table 3). Therefore, the
xenograft and unpassaged patient tumors were composed of similar
populations of phenotypically diverse cancer cell types, and in both
cases only the CD44CD24/lowLineage cells had the capacity to
proliferate to form new tumors (P 0.001).
In three of the tumors, further enrichment of tumorigenic
activity was possible by isolating the ESA subset of the
CD44CD24/low population. ESA has been used in the past to
distinguish epithelial cancer cells from benign reactive mesothelial cells (20). When ESACD44CD24/low Lineage cells were
isolated from passaged T1, as few as 200 cells consistently
formed tumors of 1 cm between 5 and 6 mo after injection,
whereas 2,000 ESACD44CD24/lowLineage cells or 20,000
CD44CD24 cells always failed to form tumors (Table 3). Ten
thousand unsorted cells formed tumors in only 3 of 12 mice. This
result suggests that the ESACD44CD24/low Lineage population was 50-fold enriched for the ability to form tumors
relative to unfractionated tumor cells (Table 3). The
ESACD44CD24/lowLineage population accounted for
24% of passage 1 T1 cells (2.55% of cancer cells). The
ESACD44CD24/lowLineage population (0.6% of cancer
cells) from unpassaged T5 cells was also enriched for tumorigenic activity, compared with ESACD44CD24/low Lineage
cells, but both the ESA and ESA fractions had some tumorigenic activity (Table 3). Among unpassaged T5 cells, as few as
1,000 ESACD44CD24/lowLineage cells consistently formed
tumors.
To determine whether the difference in tumorigenicity of the
cell populations was due to differences in cell cycle, we analyzed
these populations by flow cytometry. Comparison of the cell
cycle status of tumorigenic and nontumorigenic cancer cells
from T1 revealed that both exhibited a similar cell cycle distribution (Fig. 2). Therefore, neither population was enriched for
Fig. 3. Histology from the CD24 injection site (a; 20 objective magnification) revealed only normal mouse tissue, whereas the CD24/low injection site (b;
40 objective magnification) contained malignant cells. (c) A representative tumor in a mouse at the CD44CD24/lowLineage injection site, but not at the
CD44CD24Lineage injection site. T3 cells were stained with Papanicolaou stain and examined microscopically (100 objective). Both the nontumorigenic (d)
and tumorigenic (e) populations contained cells with a neoplastic appearance, with large nuclei and prominent nucleoli.
3986 www.pnas.orgcgidoi10.1073pnas.0530291100
Al-Hajj et al.
SEE COMMENTARY
cells at a particular stage of the cell cycle, and the nontumorigenic cells were able to undergo at least a limited number of
divisions in the xenograft model.
Six months after injection, the injection sites of 20,000 tumorigenic CD44CD24/lowLineage cells and 20,000 nontumorigenic
CD44CD24Lineage cells were examined by histology. The
CD44CD24/lowLineage injection sites contained tumors 1 cm
in diameter, whereas the CD44CD24Lineage injection sites
contained no detectable tumors (Fig. 3c). Only normal mouse
mammary tissue was seen by histology at the sites of the
CD44CD24Lineage injections (Fig. 3a), whereas the tumors
formed by CD44CD24/lowLineage cells contained malignant
cells as judged by hematoxylin and eosin-stained sections (Fig. 3b).
Even when CD44CD24Lineage injection sites from 58 mice
(each administered 1,00050,000 cells) were examined after 1629
wk, no tumors were detected. Furthermore, the tumorigenic and
nontumorigenic populations were indistinguishable morphologically. Both the tumorigenic and nontumorigenic subsets of Lineage cells from passaged and unpassaged tumors contained
95% cancer cells as judged by Wright staining or Papanicolaou
staining and microscopic analysis. By histology, the CD44CD24/low
Lineage cells and the rest of the Lineage cells had the
appearances of epithelial cancer cells (Fig. 3 d and e).
The Tumorigenic Population Is Capable of Generating the Phenotypic
Heterogeneity Found in the Initial Tumor. The ability of small
numbers of CD44CD24/lowLineage tumorigenic cells to give
rise to new tumors was reminiscent of the organogenic capacity of
normal stem cells. Normal stem cells self-renew and give rise to
phenotypically diverse cells with reduced proliferative potential. To
test whether tumorigenic breast cancer cells also exhibit these
properties, tumors arising from 200 ESACD44CD24/low
Al-Hajj et al.
Lineage T1 or 1,000 CD44CD24/lowLineage T2 cells were
dissociated and analyzed by flow cytometry. The heterogeneous
expression patterns of ESA, CD44, or CD24 in the secondary
tumors resembled the phenotypic complexity of the tumors from
which they were derived (Fig. 4 a and b vs. e and f ). Within these
secondary tumors, the CD44CD24/lowLineage cells remained
tumorigenic, whereas other populations of Lineage cancer cells
remained nontumorigenic (Table 3). Thus, tumorigenic cells gave
rise to both additional CD44CD24/lowLineage tumorigenic cells
as well as to phenotypically diverse nontumorigenic cells that
recapitulated the complexity of the primary tumors from which the
tumorigenic cells had been derived. These CD44CD24/low
Lineage tumorigenic cells from T1, T2, and T3 have now been
serially passaged through four rounds of tumor formation in mice,
yielding similar results in each passage with no evidence of decreased tumorigeneity (data not shown). These observations suggest that CD44CD24/lowLineage tumorigenic cancer cells undergo processes analogous to the self-renewal and differentiation of
normal stem cells.
Our results demonstrate that heterogeneous populations of cells
in breast cancers consist of a phenotypically distinct tumorigenic
population, as well as a much larger population that lacks this
tumorigenic potential. It is known that breast cancer cells are
genetically unstable, and thus individual breast cancer cells from the
tumorigenic population may sometimes be unable to proliferate as
a consequence of chromosomal aberrations acquired during mitosis
(2123). Nevertheless, the observation that in eight of nine tumor
specimens the tumorigenic subpopulation displayed a common
phenotype that allowed for their identification suggests that common pathways may drive this tumorigenic population.
The tumorigenic CD44CD24/lowLineage population shares
with normal stem cells the ability to proliferate extensively, and to
PNAS April 1, 2003 vol. 100 no. 7 3987
CELL BIOLOGY
Fig. 4. Phenotypic diversity in tumors arising from CD44CD24/lowLineage cells. The plots depict the CD24 and CD44 or ESA staining patterns of live human
Lineage cancer cells from T1 (a, c, and e) or T2 (b, d, and f ). T1 CD44Lineage cells (a) or T2 Lineage cells (b) were obtained from tumors that had been passaged
once in NODSCID mice. ESACD44CD24/lowLineage tumorigenic cells from T1 (c) or CD44CD24/lowLineage tumorigenic cells from T2 (d) were isolated and
injected into the breasts of NODSCID mice; e and f depict analyses of the tumors that arose from these cells. In both cases, the tumorigenic cells formed tumors
that contained phenotypically diverse cells similar to those observed in the original tumor.
patients will demonstrate clinically evident metastases. This finding
has been speculated to be due to tumor dormancy. However, an
alternative explanation consistent with a cancer stem cell model is
that cancer cells in the bone marrow of patients may arise from the
spread of either tumorigenic or nontumorigenic cancer cells, and
only when tumorigenic cells metastasize will frank tumors that are
clinically significant develop. This second explanation suggests that
the development of diagnostic reagents that allow for the prospective identification of tumorigenic cells may have prognostic significance for patients with breast cancer.
The identification of tumorigenic and nontumorigenic breast
cancer cells also has important therapeutic implications. Traditionally, drug therapies have been developed based on the ability of
these agents to cause tumor regression in animal models. Because
we have shown that the majority of cancer cells within a tumor are
nontumorigenic, therapies directed at these cells would cause tumor
regression. However, if therapies fail to target the tumorigenic cells,
then these cells would persist after therapy and be able to regenerate the tumor, resulting in tumor relapse. It is known that normal
stem cells have mechanisms that make them relatively resistant to
chemotherapy, such as increased expression of BCL-2 family proteins, increased expression of membrane transporters like breast
cancer drug resistance protein, and multiple drug resistance (29
32). The expression of such proteins in tumorigenic breast cancer
cells may make them inherently more resistant to current therapies.
The prospective identification of the tumorigenic population of
cancer cells should allow the identification of molecules expressed
in these cells that could be targeted to eliminate this crucial
population of cancer cells, leading to more effective cancer
therapies.
give rise to diverse cell types with reduced developmental or
proliferative potential (24). The extensive proliferative potential of
the tumorigenic population was demonstrated by the ability of as
few as 200 passaged or 1,000 unpassaged ESACD44CD24/low
Lineage cells to give rise to tumors (1 cm in diameter) that could
be serially transplanted in NODSCID mice. The tumorigenic
population from T1, T2, and T3 has now been purified and serially
passaged four times through NODSCID mice. This extensive
proliferative potential contrasts with the bulk of CD44 andor
CD24 cancer cells that lacked the ability to form detectable
tumors. Not only was the CD44CD24/lowLineage population of
cells able to give rise to additional tumorigenic CD44CD24/low
Lineage cells, but they were also able to give rise to phenotypically
diverse nontumorigenic cells that composed the bulk of the tumors.
This result was the case even after two rounds of serial passaging.
Thus, CD44CD24/lowLineage cells from most tumors seem to
exhibit properties of cancer stem cells. Unequivocal demonstration
of the stem cell capacity of these cells will require development of
model systems capable of tumor generation from a single cell (25).
Nonetheless, our results demonstrate that there is a hierarchy of
breast cancer cells in which some cells have the ability to proliferate
extensively, whereas the majority of tumor cells that can be derived
from this population have only limited proliferative potential in
vivo. It has previously been shown that the phenotype of acute
myelogenous leukemia leukemogenic cells is similar to that of early
hematopoietic progenitor stem cells (14). Our results suggest that
this result may also be true for tumorigenic breast cancer cells,
because early multipotent epithelial progenitor cells also have been
reported to express ESA and CD44 (2628).
The ability to separate tumorigenic and nontumorigenic populations of tumor cells should allow the molecular characterization
of these cells and elucidation of the pathways that account for their
tumorigenic potential. Furthermore, the existence of a subset of
tumorigenic cells within a tumor would provide an explanation for
a number of clinical observations in breast cancer patients. For
instance, it has been demonstrated that up to 30% of breast cancer
patients may show micrometastatic disease in their bone marrow at
the time of presentation. However, after 5 yr, only 50% of these
We thank Mark Kukaruga and Ann Marie Deslaurier for flow cytometry, Steve Ethier for tumor specimens, and Brian Clarke for the
mathematical calculation of the frequency of tumorigenic cancer cells.
This work was supported by National Cancer Institute Grant CA-075136.
Flow cytometry was supported by National Cancer Institute Grant
CA-46592.
1. Stockler, M., Wilcken, N. R., Ghersi, D. & Simes, R. J. (2000) Cancer Treat. Rev.
26, 151168.
2. Schultz, L. B. & Weber, B. L. (1999) Curr. Opin. Oncol. 11, 429434.
3. Aubele, M. & Werner, M. (1999) Anal. Cell Pathol. 19, 5358.
4. Golub, T. R. (2001) N. Engl. J. Med. 344, 601602.
5. Heppner, G. H. (1984) Cancer Res. 44, 22592265.
6. Wodinsky, I., Swiniarski, J. & Kensler, C. J. (1967) Cancer Chemother. Rep. 51,
415421.
7. Southam, C. & Brunschwig, A. (1961) Cancer 14, 971978.
8. Hamburger, A. W. & Salmon, S. E. (1977) Science 197, 461463.
9. Fialkow, P. J. (1976) Birth Defects Orig. Artic. Ser. 12, 123132.
10. Bergsagel, D. E. & Valeriote, F. A. (1968) Cancer Res. 28, 21872196.
11. Weisenthal, L. & Lippman, M. E. (1985) Cancer Treat. Rep. 69, 615632.
12. Reya, T., Morrison, S. J., Clarke, M. F. & Weissman, I. L. (2001) Nature 414,
105111.
13. Lapidot, T., Sirard, C., Vormoor, J., Murdoch, B., Hoang, T., CaceresCortes, J., Minden, M., Paterson, B., Caligiuri, M. & Dick, J. (1994)
Nature 17, 645 648.
14. Bonnet, D. & Dick, J. (1997) Nat. Med. 3, 730737.
15. Larochelle, A., Vormoor, J., Hanenberg, H., Wang, J. C., Bhatia, M., Lapidot,
T., Moritz, T., Murdoch, B., Xiao, X. L., Kato, I., et al. (1996) Nat. Med. 2,
13291337.
16. Sakakibara, T., Xu, Y., Bumpers, H., Chen, F., Bankert, R., Arredondo, M.,
Edge, S. & Repasky, E. (1996) Cancer J. Sci. Am. 2, 291300.
17. Ahrens, T., Sleeman, J., Schempp, C., Howells, N., Hofmann, M., Ponta, H.,
Herrlich, P. & Simon, J. (2001) Oncogene 20, 33993408.
18. Uchida, N., Buck, D., He, D., Reitsma, M., Masek, M., Phan, T., Tsukamoto,
A., Gage, F. & Weissman, I. L. (2000) Proc. Natl. Acad. Sci. USA 97,
1472014725.
19. Kufe, D., Nadler, L., Sargent, L., Shapiro, H., Hand, P., Austion, F., Colcher,
D. & Schlom, J. (1983) Cancer Res. 43, 851857.
20. Packeisen, J., Kaup-Franzen, C. & Knieriem, H. (1999) Hybridoma 18, 3740.
21. Murphy, K., Dennis, A. & Rosen, J. (2000) FASEB J. 14, 22912302.
22. Xu, X. (1999) Nat. Genet. 22, 3743.
23. Fukasawa, K., Weiner, F. & Mai, S. (1997) Oncogene 11, 12951302.
24. Morrison, S. J. & Weissman, I. L. (1994) Immunity 1, 661673.
25. Lagasse, E., Connors, H., Al-Dhalimy, M., Reitsma, M., Dohse, M., Osborne,
L., Wang, X., Finegold, M., Weissman, I. L. & Grompe, M. (2000) Nat. Med.
6, 12291234.
26. Stingl, J., Eaves, C., Kuusk, U. & Emerman, J. (1998) Differentiation 63, 201213.
27. Liu, A., True, L., LaTray, L., Nelson, P., Ellis, W., Vessella, R., Lange, P.,
Hood, L. & van den Engh, G. (1997) Proc. Natl. Acad. Sci. USA 94,
1070510710.
28. Gudjonsson, T., Villadsen, R., Nielsen, H. L., Ronnov-Jessen, L., Bissell, M. J.
& Petersen, O. W. (2002) Genes Dev. 16, 693706.
29. Lagasse, E. & Weissman, I. L. (1997) Cell 89, 10211031.
30. Lagasse, E. & Weissman, I. L. (1994) J. Exp. Med. 179, 10471052.
31. Domen, J., Gandy, K. L. & Weissman, I. L. (1998) Blood 91, 22722282.
32. Zhou, S., Schuetz, J. D., Bunting, K. D., Colapietro, A. M., Sampath, J., Morris,
J. J., Lagutina, I., Grosveld, G. C., Osawa, M., Nakauchi, H. & Sorrentino, B. P.
(2001) Nat. Med. 7, 10281034.
33. Porter, E. H. & Berry, R. J. (1964) Br. J. Cancer 17, 583601.
3988 www.pnas.orgcgidoi10.1073pnas.0530291100
Al-Hajj et al.
You might also like
- A Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryFrom EverandA Heartbreaking Work Of Staggering Genius: A Memoir Based on a True StoryRating: 3.5 out of 5 stars3.5/5 (231)
- The Sympathizer: A Novel (Pulitzer Prize for Fiction)From EverandThe Sympathizer: A Novel (Pulitzer Prize for Fiction)Rating: 4.5 out of 5 stars4.5/5 (119)
- Never Split the Difference: Negotiating As If Your Life Depended On ItFrom EverandNever Split the Difference: Negotiating As If Your Life Depended On ItRating: 4.5 out of 5 stars4.5/5 (838)
- Devil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaFrom EverandDevil in the Grove: Thurgood Marshall, the Groveland Boys, and the Dawn of a New AmericaRating: 4.5 out of 5 stars4.5/5 (265)
- The Little Book of Hygge: Danish Secrets to Happy LivingFrom EverandThe Little Book of Hygge: Danish Secrets to Happy LivingRating: 3.5 out of 5 stars3.5/5 (399)
- Grit: The Power of Passion and PerseveranceFrom EverandGrit: The Power of Passion and PerseveranceRating: 4 out of 5 stars4/5 (587)
- The World Is Flat 3.0: A Brief History of the Twenty-first CenturyFrom EverandThe World Is Flat 3.0: A Brief History of the Twenty-first CenturyRating: 3.5 out of 5 stars3.5/5 (2219)
- The Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeFrom EverandThe Subtle Art of Not Giving a F*ck: A Counterintuitive Approach to Living a Good LifeRating: 4 out of 5 stars4/5 (5794)
- Team of Rivals: The Political Genius of Abraham LincolnFrom EverandTeam of Rivals: The Political Genius of Abraham LincolnRating: 4.5 out of 5 stars4.5/5 (234)
- Shoe Dog: A Memoir by the Creator of NikeFrom EverandShoe Dog: A Memoir by the Creator of NikeRating: 4.5 out of 5 stars4.5/5 (537)
- The Emperor of All Maladies: A Biography of CancerFrom EverandThe Emperor of All Maladies: A Biography of CancerRating: 4.5 out of 5 stars4.5/5 (271)
- The Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreFrom EverandThe Gifts of Imperfection: Let Go of Who You Think You're Supposed to Be and Embrace Who You AreRating: 4 out of 5 stars4/5 (1090)
- Her Body and Other Parties: StoriesFrom EverandHer Body and Other Parties: StoriesRating: 4 out of 5 stars4/5 (821)
- The Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersFrom EverandThe Hard Thing About Hard Things: Building a Business When There Are No Easy AnswersRating: 4.5 out of 5 stars4.5/5 (344)
- Hidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceFrom EverandHidden Figures: The American Dream and the Untold Story of the Black Women Mathematicians Who Helped Win the Space RaceRating: 4 out of 5 stars4/5 (890)
- Elon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureFrom EverandElon Musk: Tesla, SpaceX, and the Quest for a Fantastic FutureRating: 4.5 out of 5 stars4.5/5 (474)
- The Unwinding: An Inner History of the New AmericaFrom EverandThe Unwinding: An Inner History of the New AmericaRating: 4 out of 5 stars4/5 (45)
- The Yellow House: A Memoir (2019 National Book Award Winner)From EverandThe Yellow House: A Memoir (2019 National Book Award Winner)Rating: 4 out of 5 stars4/5 (98)
- On Fire: The (Burning) Case for a Green New DealFrom EverandOn Fire: The (Burning) Case for a Green New DealRating: 4 out of 5 stars4/5 (73)
- Aamc 4RDocument76 pagesAamc 4RBellony Sanders100% (2)
- ASHRAE Std 62.1 Ventilation StandardDocument38 pagesASHRAE Std 62.1 Ventilation Standardcoolth2No ratings yet
- Collection of Blood From DonorsDocument7 pagesCollection of Blood From DonorsBernardoHernandezNo ratings yet
- Dr. LakshmayyaDocument5 pagesDr. LakshmayyanikhilbNo ratings yet
- TOFPA: A Surgical Approach To Tetralogy of Fallot With Pulmonary AtresiaDocument24 pagesTOFPA: A Surgical Approach To Tetralogy of Fallot With Pulmonary AtresiaRedmond P. Burke MD100% (1)
- FC Paed (SA) Part I Past Papers - 2011 Sept 6-4-2014Document8 pagesFC Paed (SA) Part I Past Papers - 2011 Sept 6-4-2014matenten0% (1)
- BCSC Reading Schedule 2019-2020Document11 pagesBCSC Reading Schedule 2019-2020Mari Martinez100% (1)
- Project Name: Proposed Icomc & BMC Building Complex Phe Design ReportDocument19 pagesProject Name: Proposed Icomc & BMC Building Complex Phe Design ReportAmit Kumar MishraNo ratings yet
- Summary of MetabolismDocument33 pagesSummary of Metabolismmememe123123No ratings yet
- Khan Academy Notes - CellsDocument42 pagesKhan Academy Notes - Cellsmememe123123100% (1)
- Master AAMC MCAT-2015 Topics List: Reorganized and Duplicates Removed 1. Amino AcidsDocument30 pagesMaster AAMC MCAT-2015 Topics List: Reorganized and Duplicates Removed 1. Amino AcidsSukhvir AujlaNo ratings yet
- Chapter 15 Acid Base Equilibrium and BuffersDocument9 pagesChapter 15 Acid Base Equilibrium and Buffersmememe123123No ratings yet
- Topic8Assignment CritiqueArticleDocument5 pagesTopic8Assignment CritiqueArticlearhcee11No ratings yet
- Hi Colt, I Would Like To Follow Up With You in Regards To The Seat Cushion Frame Support Wire. Thank YouDocument1 pageHi Colt, I Would Like To Follow Up With You in Regards To The Seat Cushion Frame Support Wire. Thank Youmememe123123No ratings yet
- Cancer Bio Exam Article Supplemental InfoDocument8 pagesCancer Bio Exam Article Supplemental Infomememe123123No ratings yet
- Ochem 5 Lab TechniquesDocument50 pagesOchem 5 Lab Techniquesmememe123123No ratings yet
- Leitz & Kavalali 2011Document9 pagesLeitz & Kavalali 2011mememe123123No ratings yet
- Evidence Supporting Presence of Two Pacemakers in Rat ColonDocument12 pagesEvidence Supporting Presence of Two Pacemakers in Rat Colonmememe123123No ratings yet
- Master AAMC MCAT-2015 Topics List: Reorganized and Duplicates Removed 1. Amino AcidsDocument30 pagesMaster AAMC MCAT-2015 Topics List: Reorganized and Duplicates Removed 1. Amino AcidsSukhvir AujlaNo ratings yet
- What's On The MCAT 2015Document128 pagesWhat's On The MCAT 2015kape1oneNo ratings yet
- Microbial GeneticsDocument11 pagesMicrobial GeneticsJennie M. MagdirilaNo ratings yet
- MCAT 2015 OutlineDocument128 pagesMCAT 2015 OutlinequirkyheadbandsNo ratings yet
- MCAT Study PlanDocument55 pagesMCAT Study Planmememe123123No ratings yet
- 10 Alkenes ReactionsDocument1 page10 Alkenes ReactionsRakesh Kumar SutharNo ratings yet
- Chem 242 Presentation SummaryDocument2 pagesChem 242 Presentation Summarymememe123123No ratings yet
- 10 Alkenes ReactionsDocument1 page10 Alkenes ReactionsRakesh Kumar SutharNo ratings yet
- C242f14sec1syllabus PDFDocument20 pagesC242f14sec1syllabus PDFmememe123123No ratings yet
- Reference FormDocument2 pagesReference Formmememe123123No ratings yet
- Khan Academy Visual Perception GuideDocument1 pageKhan Academy Visual Perception Guidemememe123123No ratings yet
- C 242 F 14 T 1Document15 pagesC 242 F 14 T 1mememe123123No ratings yet
- Saudi Board Exam - Surgery Questions from 2009 Exam (192/200Document47 pagesSaudi Board Exam - Surgery Questions from 2009 Exam (192/200Rahmah Shah Bahai100% (2)
- UntitledDocument221 pagesUntitledlaljadeff12No ratings yet
- Angina Pectoris: Causes, Symptoms and TreatmentsDocument17 pagesAngina Pectoris: Causes, Symptoms and TreatmentsGaming ModeNo ratings yet
- Beta Lactam Antibiotics Structure, Classification and MechanismDocument15 pagesBeta Lactam Antibiotics Structure, Classification and MechanismNiharika ModiNo ratings yet
- Kerry Washington. Family Secret.Document3 pagesKerry Washington. Family Secret.yulya.shevchenko110No ratings yet
- Psihogeni Neepileptički Napadi Kao Dijagnostički Problem: AutoriDocument4 pagesPsihogeni Neepileptički Napadi Kao Dijagnostički Problem: AutorifhdhNo ratings yet
- Phenol Hazards and PrecautionsDocument3 pagesPhenol Hazards and PrecautionsMarnel Roy MayorNo ratings yet
- Which Is More Effective in Treating Chronic Stable Angina, Trimetazidine or Diltiazem?Document5 pagesWhich Is More Effective in Treating Chronic Stable Angina, Trimetazidine or Diltiazem?Lemuel Glenn BautistaNo ratings yet
- Atls 1Document5 pagesAtls 1Mega MalyndaNo ratings yet
- Nursing Audit ScheduleDocument2 pagesNursing Audit ScheduleArvinjohn Gacutan0% (1)
- Botany (Virus)Document48 pagesBotany (Virus)Madhuri DeviNo ratings yet
- Curiculum Vitae Iyere Faith-1Document5 pagesCuriculum Vitae Iyere Faith-1Halleluyah HalleluyahNo ratings yet
- F&F LetterDocument3 pagesF&F LetterJaskeerat Singh50% (2)
- eBR PharmaDocument5 pageseBR PharmaDiana OldaniNo ratings yet
- Run TroopDocument65 pagesRun TroopRob Colares100% (2)
- Sickle-Cell AnemiaDocument11 pagesSickle-Cell Anemiahalzen_joyNo ratings yet
- Low Back Pain Dr. Hardhi PRanataDocument57 pagesLow Back Pain Dr. Hardhi PRanataPerwita ArumingtyasNo ratings yet
- Vector and Pest Control in DisastersDocument10 pagesVector and Pest Control in DisastersTaufik RizkiandiNo ratings yet
- The Nature of Feeding and Swallowing Difficulties in PDFDocument118 pagesThe Nature of Feeding and Swallowing Difficulties in PDFLisa NurhasanahNo ratings yet
- Beddtiot ORTHO / Orthodontic Courses by Indian Dental AcademyDocument60 pagesBeddtiot ORTHO / Orthodontic Courses by Indian Dental Academyindian dental academyNo ratings yet
- Division Memorandum - s2020 - 241Document16 pagesDivision Memorandum - s2020 - 241ARLENE MARASIGANNo ratings yet
- What Is Drug ReflectionDocument8 pagesWhat Is Drug ReflectionCeilo TrondilloNo ratings yet
- From MaureenDocument1 pageFrom MaureenWKYC.comNo ratings yet