Professional Documents
Culture Documents
Protein Purification
Uploaded by
Raja Mohan GopalakrishnanCopyright
Available Formats
Share this document
Did you find this document useful?
Is this content inappropriate?
Report this DocumentCopyright:
Available Formats
Protein Purification
Uploaded by
Raja Mohan GopalakrishnanCopyright:
Available Formats
26/09/2016
ProteinPurification
ProteinPurification
CarolineRitchie(carolinedotritchie103atgmaildotcom)
IowaStateUniversity,UnitedStates
DOI http://dx.doi.org/10.13070/mm.en.2.134
Date lastmodified:20150902originalversion:20121117
Citeas MATERMETHODS20122:134
Abstract
Anindepthreviewofcolumnchromatographyforproteinpurificationandsurveyresultfrom305
formalpublications.
Purified proteins are required for many experimental applications, including structural studies and in
vitro biochemical assays. Proteins can be obtained from tissue or, more often, by their overexpression in a
modelorganism,suchasbacteria,yeast,ormammaliancellsinculture.Proteinpurificationinvolvesisolating
proteins from the source, based on differences in their physical properties. The objective of a protein
purification scheme is to retain the largest amount of functional protein with fewest contaminants. The
purificationschemeofaproteinmustbeoptimizedtocompletethisprocessintheleastnumberofsteps.
The
article
reviews
four
types
of
column
chromatography that are commonly used in protein
purification, and discusses the advantages,
disadvantages, and potential problems of each. This
article also reports a Labome survey on 98
publications. The survey indicates that affinity column
chromatography, mainly that based on HIS, GST,
andFLAGtags,andsizeexclusionchromatographyare
the main methods cited in the publications. GE
Healthcare is the major supplier of reagents and
instrumentsusedinproteinpurfication.
Figure1.ATrisbufferedsolutioncontainsTris
baseanditsconjugateacid.ThepKaofTrisat
25Cis8.06,indicatingthatatpH=8.06,50%of
theTrisisprotonated(initsacidicform)and50%
isdeprotonated(initsbasicform).
DevelopingaProteinPurificationScheme
The most important consideration in the development of a protein purification scheme is the downstream
applicationofthepurifiedprotein.Boththequantityandpurityoftheproteinmustbesufficientforexperimental
analysis.Additionally,informationaboutthebehavioroftheproteinmustbetakenintoconsideration,aswell
foldedandfunctionalproteinisrequiredfordownstreamstudies.Duringpurificationandsubsequentstorage,
manyprocessescanoccurthataffectproteinquality:proteinunfolding,aggregation,degradation,andlossof
function.Carefulplanningtopurifyproteinasquicklyaspossibleandunderthemoststabilizingconditionswill
maximizethechanceofasuccessfulpurificationscheme.
BufferingComponent
Thesolutionconditionsofaproteinateachstepofthepurificationschemeareessentialinmaintainingprotein
stabilityandfunction.ProteinsshouldbekeptinawellbufferedenvironmenttopreventsuddenchangesinpH
thatcouldirreversiblyaffecttheirfolding,solubility,andfunction.
https://www.labome.com/method/ProteinPurification.html
1/16
26/09/2016
ProteinPurification
Abufferisasolutioncontainingaconjugateacid/basepair.ThepHrangeofabufferis
basedonitspKa,definedasthepHatwhich50%ofthemoleculesareintheiracidic
formand50%areintheirbasicform(Figure1).Ageneralruleregardingbuffersisthat
thepHofthebuffersolutionshouldbewithin1.0pHunitofthepKainordertoprovide
appropriate buffering capacity. This ensures that there is a sufficient amount of the
molecule in both its acidic and basic forms to neutralize the solution in case of H+ or
Figure2.An
Aktaprimeplus
systemforthe
automated
chromatographic
separationof
proteins.From
GE.
OH influx. Thus, buffers prevent pH changes that could negatively affect protein
stability.
A good buffer must exhibit the
followingcharacteristics[1]:
1.Watersolubility
2.Chemicalstability
3.Highbufferingcapacityat
desiredpH
4.Compatibilitywith
analyticaland
experimentalapplications
5.Compatibilitywithother
solutioncomponents
Manycomponentscanserveasbiologicalbuffers.The
most commonly used buffering components have a
nearneutralpKa,astheycanbeusedataphysiological
pH. Four of the most common biological buffers are
listedinTable1,alongwiththepHrangeatwhichthey
Figure3.NiNTAligandcovalentlyattachedtoa
crosslinkedagarosematrixfortheselective
purificationofpolyhistidinetaggedproteins.
HistidineresiduescancoordinatetotheNi2+ion
byreplacingtheboundwatermolecules
(indicatedbyredarrows).Imidazoleorfree
histidinecanthenbeusedtoelutetheprotein,
throughtheirabilitytocoordinatewiththeNi2+ion
anddisplacetheboundprotein.FromBioKe.
can be used, and advantages and disadvantages that
mightaffecttheirusageinproteinpurification.Typically,thesebuffersareusedatconcentrationsabove25mM
toensureadequatebufferingcapacity.
SolutionAdditives
Buffer
pHrange
Advantagesand
Disadvantages
pHisnot
dependenton
temperature
Inexpensive
Phosphate
5.88.0
Transparentin
UVrange
Cannotbeused
withdivalent
cations
https://www.labome.com/method/ProteinPurification.html
Inadditiontoanappropriatebufferingsystem,solutions
used during protein purification from lysis to storage
often contain many other components that play a role
infacilitatingproteinpurity,stability,andfunction.
Protease inhibitors are often added to the lysis buffer
andinearlystepsofthepurificationschemetoprevent
degradation of the target protein by endogenous
proteases. These are generally not needed toward
later stages of the purification, as most or all of the
contaminatingproteaseshavebeenseparatedfromthe
protein of interest. Metal chelating reagents, such as
EDTA or EGTA, are often added to the storage buffer.
2/16
26/09/2016
ProteinPurification
Cannotbe
autoclaved
pHissomewhat
MOPS
6.57.9
dependenton
temperature
Highbuffering
ThesemetalchelatorsbindtoMg2+and,thus,prevent
cleavage of the purified protein by contaminating
metalloproteases. Other additives are often used to
protect proteins against damage and enhance their
solubility.
capacityat
Additives should only be used if necessary. Trial and
error is often required to determine the specific
physiologicalpH
additives that are beneficial to a particular protein
purificationscheme.
Cannotbe
autoclaved
HEPES
6.88.2
Otherfactorstoconsider
pHissomewhat
dependenton
Otherfactorsalsocontributetoproteinstabilityduringa
temperature
Canformradicals
during its purification is always the best. Designing a
purification scheme that uses the minimum number of
undercertain
steps in the shortest amount of time ensures the
conditions[2]
highestyieldoffunctionalprotein.Additionally,itisoften
purificationscheme.Theleastmanipulationofaprotein
best to keep the protein cold throughout the
pHisdependent
ontemperature
anddilution
Inexpensive
Tris
7.59.0
Caninterferewith
activityofsome
enzymes[3],[4]
Transparentin
UVrange
Table1.Mostcommonlyusedbiologicalbuffers
forproteinpurification.Buffersmaintaintheir
bufferingcapacitywithinaspecificpHrange,and
characteristicsofsomebufferingcomponents
couldinterferewithparticularchromatographic
proceduresoranalysis.Summarizedfrom
Promega,SigmaAldrich,Applichem,Embl.de.
purification. Typically, purification is performed at 4C,
as this lower temperature both slows down the rate of
proteolysis (in the event of contaminating proteases)
andpromotesstructuralintegrityofproteins.
IntroductiontoColumnChromatography
EquipmentforColumnChromatography
Theprincipleofcolumnchromatographyistoseparate
alargepoolofproteinsintomanysmallerpools,some
of which are enriched in the protein of interest. While
expensive and specialized equipment is available for
column chromatography, only basic equipment is
required.
Basicequipmentforcolumnchromatography:
1.Stationaryphase:aninert
matrix,oftenwithan
attachedfunctionalgroup
Type
Function
tofacilitateprotein
interaction,usedto
separateproteins.The
choiceofstationary
Protectagainstoxidative
ReducingAgents
phaseandfunctional
damage
groupdependsonboth
thetypeof
chromatographythatis
https://www.labome.com/method/ProteinPurification.html
CommonlyUsedReagents
2mercaptoethanol
(BME)
Dithiothreiotol(DTT)
Tris(2carboxyethyl)
phosphine(TCEP)
3/16
26/09/2016
ProteinPurification
beingperformedandthe
methodbywhichitwillbe
carriedout.
2.Column:acylindrical,
Inhibitendogenous
glassreservoiravailable
ProteaseInhibitors
proteasesfrom
invariouslengthsand
degradingproteins
diameters.Columnscan
bepurchasedprepacked
withastationaryphase
andreadytoattachto
automated
chromatographysystems
Inactivate
MetalChelators
(discussedinPart2of
metalloproteases
thissection)orcanbe
boughtemptyformanual
packing.Differenttypesof
columnsarerequiredfor
automatedversusgravity
Stabilizeproteinstructure
Osmolytesflowchromatography.
andenhancesolubility
Leupeptin(serineand
cysteineprotease
inhibitor)
PepstatinA(asparticacid
proteaseinhibitor)
PMSF(serineprotease
inhibitor)
EDTA
EGTA
Glycerol
Detergents(e.g.,CHAPS,
NP40,TritonX100)
3.Solvents:buffers
Sugars(e.g.,glucose,
containingadditivesused
sucrose)
forequilibrating,washing,
andelutingproteinsfrom
Salts(e.g.,NaCl,KCl,(NH4 )2 SO4
IonicStabilizers
Enhancesolubility
thestationaryphase.
Differenttypesof
chromatographyrequire
Table2.Additivescommonlyusedinproteinpurificationbuffersto
differentsolvent
increasestabilityofproteins.SummarizedfromThermoFisherPierceand
Embl.de. conditions.
4.Collectiontubes:vessels
forcollectingelution
samples.Specialtubes
arerequiredfor
automatedfraction
collectorshowever,any
tubeorvesselis
appropriateformanual
fractioncollection.
Figure4.Schematicofaffinity
purificationusingProteinA,G,or
L.A.Antibodiescontainseveral
regionsofinterest:Fc(fragment,
constant)thatisthesameforall
https://www.labome.com/method/ProteinPurification.html
5.Assaytomeasurepurity:
anapproachfor
determiningtherelative
amountofaspecific
proteintothetotalprotein
inasample.Thefractions
containingtheproteinof
interestmustbe
determinedaftereach
stepbeforeproceedingto
thenextstepinthe
purificationscheme.
Somecommonmethods
ofanalyzingpuritywillbe
discussedinalater
section.
SystemsforColumnChromatography
4/16
26/09/2016
ProteinPurification
proteinsofthesameclass,Fv
(fragment,variable)thatoffers
specificityforeachantibody,and
Fab(fragment,antigenbinding)
thatactuallycontactsthespecific
antigen.B.Specificclassesof
antibodiescanbenonselectively
purifiedthroughaffinitypurification
inwhichtheligand(ProteinA,G,
orL)isconjugatedtoamatrix.C.
ProteinA,G,orLcanalsobe
usedtopurifyspecificproteins.In
thiscasetheantibodywillserve
asanintermediateligand
providingselectivityforitsantigen.
Column chromatography can be performed with automated systems
(Figure2),whichuseapumptoforcesolventoverapackedcolumnat
a set flow rate, or can be run by gravity flow. Both automated and
gravity flow systems can be coupled to automatic fraction collecting
systems. There are advantages and disadvantages to each system.
GEHealthcareAKTAFPLCsystemisaverycommonchoice[514].
TypesofColumnChromatography
The four major types of column chromatography include affinity
chromatography, ion exchange chromatography (IEX), hydrophobic
interactionchromatography(HIC),andsizeexclusionchromatography
(SEC).Mostpurificationschemesrequiretheuseofmorethanoneof
these
types
of
chromatographic procedures
TypeofSystem
Advantages
to yield the necessary purity
for downstream applications.
Choosing
the
most
Disadvantages
Canletrunbyitself
Oftencoupledtoan
absorbancedetector
Automated
Canprogram
equilibrationand
washsteps
Easytosetupa
gradientforelution
Requiresspecific,
costlyequipment
Maximumflowrate
isdependenton
these methods is essential in
optimizing
a
protein
purificationscheme.
pressurelimitof
column
By analyzing the sequence of
a
protein,
unique
characteristics
can
be
identified that might assist in
Veryreproducible
Lessexpensive
Theuserhasmore
control
GravityFlow
Canmake
adjustmentsduring
run
appropriate chromatographic
method(s) and the order of
Morelaborintensive
Flowroteislimited
bygravity
its purification. The size and
charge of a protein (at a
specific
pH)
can
be
determined, along with the
identificationoflargestretches
ofhydrophobicresidues.
AffinityChromatography
Table3.Systemsforthechromatographicseparationofproteins.
SummarizedfromGE.
Affinity chromatography relies
on the specific and reversible
binding of a protein to a
matrixbound ligand. The
Typeof
Chromatography
ligand can either bind directly
to the protein of interest or to
a tag that is covalently
attachedtotheprotein.Affinity
chromatography is often the
most
robust
Affinity
purification
https://www.labome.com/method/ProteinPurification.html
Separates
ProteinsBy
Aspecific
interaction
BindWith
Nocompeting
ligand
EluteWith
Competingligand
(specific)
conditionsthat
disrupt
protein/protein
5/16
26/09/2016
ProteinPurification
interactions(non
specific)
procedure and is typically
used in early stages of the
purification
scheme.
Depending
downstream
on
the
application,
affinity purification might be
theonlychromatographicstep
required to achieve adequate
purity.
The stationary phase for
affinity chromatography is
IonExchange
Netsurfacecharge
Lowionicstrength
Highionicstrength
Increased(cation
exchange)or
decreased(anion
exchange)pH
Hydrophobic
Interaction
Hydrophobicity
Highionicstrength
Lowionicstrength
SizeExclusion
Hydrodynamic
radius
Table4.Thefourmostcommontypesofcolumnchromatographyusedin
proteinpurification.
made of an inert matrix
covalentlyattachedtoaligandthatspecificallybindsto
a protein or group of proteins. The inert matrix is
typically composed of crosslinked agarose or
polyacrylamide. Proteins can be purified by affinity
chromatographyinaselectiveornonselectivemanner.
Inselectiveaffinitychromatography,aligandspecificfor
a protein or a covalently attached tag is used. In non
selectiveaffinitychromatographysuchasProteinA,G,
L for immunoglobulin, or heparin for DNAbinding
proteins, or lectin for glycoproteins, the ligand binds to
agroupofproteinswithsimilarbindingcapabilities.
In both types of affinity chromatography, proteins are
loaded on the column under conditions that influence
bindingbetweentheprotein(ortag)anditsligand.The
bound protein is washed under conditions that do not
disruptthespecificinteraction,butthatcandisruptany
nonspecific interactions between contaminating
Figure5.Thenetchargeonaproteinis
influencedbythepHofitssolvent.AtpH=pI,the
proteinhaszeronetchargeand,therefore,will
notbindtoacationexchangeorananion
exchangestationaryphase.AdjustingthepH
aboveorbelowthepIoftheproteinwillleadtoa
netcharge,andproteinbindingtoeitherananion
exchange(pH>pI)oracationexchange(pH<
pI)stationaryphase.
proteins and the stationary phase. The bound protein is then eluted with a buffer containing a competing
molecule or conditions that disrupt all protein/protein interactions. Competing molecules bind to the ligand,
displacing the protein of interest. This competing molecule is typically removed from the protein of interest
eitherthroughanotherchromatographicprocedureordialysis.Methodsforelutingproteinsfromthestationary
phase by disrupting all protein/protein interactions include adjusting the pH or ionic strength of the buffer.
These methods can affect protein stability, and it is suggested that the eluted protein be immediately
neutralized or diluted to minimize protein damage. For some forms of affinity chromatography, alternative
elutionconditionshavebeendescribedtomaximizetheyieldoffunctionalprotein[15,16].
Proteinto
Purify
Antibody
(antigen
specific)
Polyhistidine
Ligand
Antigenic
peptide
EluteWith
Indesigningaplasmidforproteinexpression,affinity
tags can be inserted on either the N or Cterminus
Freepeptide
known structures) to assist in purification. See
Labome survey on protein/protide tags for a list of
Imidazoleor
https://www.labome.com/method/ProteinPurification.html
(or in rare cases, in flexible loops of proteins with
commontagsandtagrelateddiscussions.
6/16
26/09/2016
ProteinPurification
taggedprotein
Ni2+orCo2+
freehistidine
FLAGtagged
protein
FLAG
specific
antibody
FLAG
peptideor
lowpH
specific interaction that occurs between an antibody
and its antigen (the sequence recognized by the
GSTtagged
protein
Reduced
glutathione
Free
glutathione
coupled to a matrix for specific binding of the
Myctagged
protein
Mycspecific
antibody
LowpH
Antibody
(classspecific)
ProteinA,
G,orL
Extremesin
pH
DNAbinding
protein
Heparin
Highionic
strength
Antibodies are often purified based on the highly
antibody). A peptide containing the antigen can be
antibody. Lowering the pH of the elution buffer
disrupts the antibody/peptide interaction to release
the bound antibody. This method is often used
commerciallyintheisolationofantibodiesfromcrude
serum.
Proteins
can also
Table5.Examplesofselectiveandnonselective
formsofaffinitychromatographywiththefunctional
group(ligand)usedforspecificityandtypical
elutionconditions.SummarizedfromThermo
FisherPierceandGE.Seebelowforthecommon
suppliersfordifferenttypesofcolumnsbasedona
Labomesurveyofover200publications.
beaffinity
purifiedin
a
non
selective
manner.
In non
selective
Figure6.Ionexchange
chromatography.Proteinsarebound
toachargedstationaryphaseatlow
ionicstrength.Theboundproteins
canbeelutedeitherbyincreasing
theionicstrengthofthebufferorby
adjustingthepH.
purification, the ligand attached to the stationary phase binds to a group of proteins with similar binding
partners.AnexampleofnonselectiveaffinitypurificationisinthepurificationofDNAbindingproteins.Heparin
mimics DNA both in its structure and its charge, and can be used as the ligand for the affinity purification of
DNAbinding proteins. While all DNAbinding proteins could theoretically bind to this stationary phase, most
other proteins will flow through without binding, leading to sufficient enrichment of the protein of interest.
Another example is the enrichment of antibodies by the binding of their constant (Fc) region to the ligand,
ProteinA,G,orL.GEHealthcareproteinAaffinitycolumnsareacommonchoice[1719],asisproteinG[20].
Usingasimilarmodeofbinding(anantibodyboundtoaProteinA,G,orLligand),theantigenbinding(Fab)
regionoftheantibodyisstillavailableforbindingtoitsspecificantigen.Thus,specificproteinscanbepurified
basedonthespecificinteractionbetweenthecoupledligand/antibodyandtheantibody/antigen.
CommonproblemsandtroubleshootingarelistedinTable6.
https://www.labome.com/method/ProteinPurification.html
7/16
26/09/2016
ProteinPurification
IonExchangeChromatography
(IEX)
Problem
Proteindoesnotbind
charge, through electrostatic
interactions
that
occur
between
proteins
and
charged stationary phase.
Two types of IEX exist: (1)
anion exchange (positively
Proteindoesnotelute
charged stationary phase that
binds to negatively charged
proteins)
and
(2)
cation
exchange (negatively charged
stationary phase that binds to
positively charged proteins).
Ionexchangechromatography
Lowresolution
is commonly used as an
intermediate step in a protein
purification scheme however,
it can yield high resolution for
some proteins when used
earlier or later during the
purification.
All proteins exhibit a net
Solution
Checkplasmidsequence
Tagwasnottranslatedoris
ormovetagtoadifferent
notaccessible
location
Ionexchangechromatography
(IEX) separates proteins
based on their net surface
Cause
Bindingconditionsarenot
appropriate
Adjustbufferconditions
Notenoughtimewas
allowedforbinding
Decreaseflowrateorstop
columntoallowincubation
Affinitybetweenligandand
tagisveryhigh
Increaseconcentrationof
competitor(specific)or
stringencyofconditions
(nonspecific)
Proteinaggregatedon
column
Adjustbufferconditionsfor
moreproteinstability
Flowrateiseithertoofast
ortooslow
Adjustflowrate
Columnwasnotwashed
sufficiently
Washwithhigher
stringencybufferClean
stationaryphaseaccording
tomanufacturer
Proteinaggregatedon
column
Adjustbufferconditionsfor
moreproteinstability
Elutionconditionsarenot
stringentenough
Increaseconcentrationof
competitor(specific)or
adjustconditions(non
specific)
Proteinisunfoldedor
aggregated
Adjustbufferconditionsfor
moreproteinstability
Proteinlosesactivityduring
Acofactorrequiredfor
procedure
activitywasremoved
duringpurification
Addcofactor
charge that depends on the
aminoacidcompositionofthe
protein and any covalently
Table6.Troubleshootingaffinitychromatography.SummarizedfromGE.
attachedmodifications.ThenetchargeofaproteinisinfluencedbythepHofthesolventthatitisdissolvedin,
assolventsexchangehydrogenionswithproteins.Theisoelectricpoint(pI)ofaproteinisthepHatwhichthe
proteinhasnonetcharge.AtapHabovethepI,aproteinwillhaveanetnegativecharge,whileapHbelow
thepIwillleadtoanetpositivecharge.Thus,thepHofthesolventcanbeadjustedtofacilitatebindingtoIEX
ortopromoteelutionofaboundprotein.
Theoretically, all proteins can bind to both cation exchange and anion exchange if the pH of the buffer is
adjusted accordingly. However, for protein purification, the stability of the protein is the most important
consideration in choosing purification conditions and, thus, the most appropriate column for protein binding.
Therefore,itisnecessarytodeterminewhetherconditionsrequiredforbindingtoeitherformofionexchange
chromatography affect protein stability and function. Typically, conditions for binding to one type of ion
exchangeismoresuitableforaparticularproteinthantheother.
Knowledge of a protein's isoelectric point can aid in determining the most appropriate type of ion exchange
chromatography. Online tools are available for calculating the theoretical pI of a protein
EXPASY
. These
calculationsarebasedentirelyontheaminoacidsequenceofaproteinanddonottakethethreedimensional
https://www.labome.com/method/ProteinPurification.html
8/16
26/09/2016
ProteinPurification
structure of the protein into consideration. In its native state, some
residues of a protein are more exposed than others and, thus, the
actual pI and net surface charge is sometimes different from those
determinedtheoretically[21].Thedistributionofaminoacidsrelative
to one another can also affect a protein's pI [22], and this is not
taken into consideration with theoretical pI calculations. Some
techniquesallowfortheexperimentaldeterminationofaprotein'spI
initsnativestate,includingelectrophoreticisoelectricfocusing[23],
capillary isoelectric focusing [24], and a more recent high
throughputluminescencebasedapproach[25].
Typically,bindingofaproteintoIEXmustbedeterminedbytrialand
error, using solvents with a range of pH values, to determine the
optimumpHforproteinretention.AsolventpHthatisaboutonepH
unit away from the pI is usually sufficient for protein binding [26]
however,insomecasesapHfurtherfromthepIisrequired[27].
ThestationaryphaseofIEXiscomposedofaninertagarosebased
or polymeric matrix with a covalently bound charged group. Matrix
particlesareavailableinmanysizes,andcaneitherbenonporous
orcancontainporesofvariablesize.Thechoiceofmatrixdepends
on the desired binding capacity, resolution, and flow rate. Smaller
particlesizesofferhigherresolution,butrequirelowerflowratesand
take longer to run. Porous matrices offer a higher binding capacity
than nonporous matrices. In a nonporous matrix, proteins cannot
enter the resin and, thus, offer increased sample recovery, greater
resolution, and shorter run times than porous matrices. One study
showed that, for most proteins, resolution and recovery are similar
when separated by a porous versus nonporous matrix however,
Figure7.Hydrophobicinteraction
chromatography.Athighionic
strength,proteinsarepartially
desolvated,causingthemtoadopt
alternateconformationsinwhich
normallyburiedhydrophobic
residuesaremoreexposed.These
residuescanthenformhydrophobic
interactionswiththehydrophobic
functionalgroupsconjugatedtoa
matrix.Loweringtheionicstrength
causestheproteintorefoldintoits
nativeconformation,buryingits
hydrophobicresidues.This
decreaseshydrophobicinteractions
betweentheproteinandstationary
phase,facilitatingproteinelution.
forlargerproteins,porousmatricescausealossinresolutionbasedonsizeexclusioneffects[28].
The four charged functional groups most commonly used in
IEXareshownbelow.Theseareclassifiedaseitherstrongor
weakexchangers,withstrongexchangersbeingionizedovera
widerpHrangethanweakexchangers.
Anionexchange(positivelychargedstationaryphase):
1.quaternaryammonium(Q)stronganionexchanger
2.diethylaminoethyl(DEAE)weakanionexchanger
Cationexchange(negativelychargedstationaryphase):
1.methylsulfonate(S)strongcationexchanger
Figure8.Proteinelutedfroma
hydrophobicinteractioncolumnwitha
decreasingsalt(ammoniumsulfate)
https://www.labome.com/method/ProteinPurification.html
2.carboxymethyl(CM)weakcationexchanger
9/16
26/09/2016
ProteinPurification
gradient.Fractionswereanalyzedforboth
totalproteincontentandactivityspecific
totheproteinofinterest.Thepeak
centeredatfraction45containsthe
proteinofinterest,asindicatedbyprotein
activity.From
http://www.insectscience.org/9.04/.
Strong exchangers should be used when the pH required for
binding is very acidic or basic (assuming this pH is also
appropriate for maintaining protein stability), as the functional
groups remain charged over a larger pH range, [29]. Weak
exchangers have been shown to provide less retention of
some proteins, enabling their elution at lower ionic strength
[30]. Because high ionic strength can affect the stability of
someproteins[31],weakexchangersmightbemoresuitableforproteinsthatdonotrequireanextremepHfor
binding.
Inionexchangechromatography,thestationaryphaseisfirstequilibratedwithalowionicstrengthbuffer.The
protein sample is then loaded onto the stationary phase in the same low ionic strength buffer used for
equilibration.Theboundproteiniswashedextensivelybeforeelutionwithahigherionicstrengthbufferor,in
some cases, a buffer with altered pH (Figure 6). Counterions in the elution buffer interact with the charged
stationary phase, displacing the bound protein. Certain salts are more efficient for use in ion exchange
chromatographythanothers,duetotheirabilitytodisplaceboundproteinsandtheireffectsonproteinstability
[32].TypicallyNaClorKClisusedforelution,withNa+orK+servingasthecountercationincationexchange
chromatographyandClservingasthecounteranioninanionexchangechromatography.Alternatively,thepH
ofthebuffercanbealteredtoreducetheprotein'schargeanddisruptitsinteractionwiththestationaryphase.
Forproteinsboundtoacationexchangestationaryphase,increasingthebufferpHwillleadtoalesserpositive
charge on the protein, and subsequent elution from the column. For proteins bound to an anion exchange
stationaryphase,decreasingthebufferpHwillleadtoalessernegativechargeontheprotein,andsubsequent
elutionfromthecolumn.
Alternatively, the pH of the buffer can be adjusted so
that the protein of interest does not bind to the ion
exchange stationary phase while contaminating
proteins do. In this case, the protein of interest is
collectedintheflowthrough,whilesomecontaminating
proteinsareremovedbytheirbindingtothestationary
phase.
As with other chromatographic methods, ion exchange
chromatography may require some troubleshooting to
determine the optimum conditions for protein binding,
proteinelution,andadequateresolution.
HydrophobicInteractionChromatography(HIC)
Hydrophobic interaction chromatography (HIC)
separates proteins based on their hydrophobicity, and
is often used as an intermediate step in a purification
scheme.Proteinsareboundtoastationaryphaseina
Figure9.Amixtureofthreeproteinsofvarying
hydrodynamicradiusisloadedontoasize
exclusioncolumn.Largeproteinselutefirst,as
theyareunabletoentertheporesofthematrix
andhaveastraightforwardpaththroughthe
column.Smallerproteinscanenterthepores,
haveamoreconvolutedpathand,thus,take
longertotraversethematrixandelutefromthe
column.
high ionic strength buffer and,
Problem
Cause
Solution
thus, HIC can typically be
performed immediately after
Runmoreequilibration
https://www.labome.com/method/ProteinPurification.html
10/16
26/09/2016
Proteindoesnotbind
ProteinPurification
Columnwasnot
equilibrated
bufferthroughcolumnand
reloadprotein
ionexchangechromatography
Ionicstrengthofbinding
bufferistoohigh
Lowerionicstrengthof
buffer
AdjustbufferpH(lowerfor
cationexchange,higherfor
anionexchange)
dilution required. HIC is also
commonly performed after an
pHisnotfarenoughfrom
pI
Proteindoesnotelute
Lowresolution
with no buffer exchange or
ammonium
sulfate
precipitation, a procedure that
canbeusedtoquicklyremove
Ionicstrengthofelution
bufferistoolow
Increaseionicstrength
Proteinaggregatedon
column
Adjustbufferconditionsfor
moreproteinstability
Flowrateiseithertoofast
ortooslow
Adjustflowrate
Columnwasnotwashed
sufficiently
Washwithahigherionic
strengthbufferClean
stationaryphaseaccording
tomanufacturer
Proteinaggregatedon
column
Adjustbufferconditionsfor
moreproteinstability
The presence of salt ions in
Proteinisunfoldedor
aggregated
Adjustbufferconditionsfor
moreproteinstability
solution can lead to partial
unfolding of a protein and the
Proteinlosesactivityduring
Acofactorrequiredfor
procedure
activitywasremoved
duringpurification
proteinsbyprecipitatingsome,
but not all, proteins with salt.
HICissometimesapplicablein
early steps of a purification
scheme or as a final step in
theremovaloftraceimpurities
fromtheproteinofinterest.
exposure of normally buried
Addcofactor
Table7.Troubleshootingionexchangechromatography.Summarized
fromGE.
hydrophobic
residues.
Proteins that bind to the
stationary phase adopt their
native conformation as buffer
with a lower ionic strength is
added. This decreases the
exposure of hydrophobic residues that can interact with the stationary phase, and facilitates elution from the
column.Forproteinsthatcanrefoldspontaneouslythroughadecreaseinionicstrength,HICcanbeavaluable
chromatographicmethodforpurification.
Typically, the ionic strength of the binding buffer should be as low as possible to bind the protein of interest,
while preventing its precipitation. If the ionic strength required for protein binding is high and causes
precipitation of the protein of interest, a lower ionic strength can be used. In this case, the chromatography
procedure can be used to separate all of the binding proteins from the protein of interest that flows through
withoutbinding.
Prior to loading the sample on the column, the stationary phase must be equilibrated with the high ionic
strengthbuffer(thesamebufferthattheproteinsamplewillbeappliedin).Thesampleisthenloadedontothe
column,thecolumniswashedextensively,andtheproteiniselutedwithalowionicstrengthbuffer(Figures7
and8).
Thestationaryphaseusedinhydrophobicinteractionchromatographyiscomposedofabasematrixofeither
crosslinked agarose or a synthetic copolymer. An alkyl or aryl ligand is then conjugated to the base matrix,
providingspecificityforhydrophobicmolecules.
Typesoffunctionalgroup:
https://www.labome.com/method/ProteinPurification.html
11/16
26/09/2016
ProteinPurification
1.Alkylahydrocarbonchainofvariouslengthoftenabutyloroctylgroupisused.Thebindingcapacityof
thestationaryphaseisincreasedwithincreasingalkylchainlength[33].Thefunctionalgroupsbind
proteinsbasedentirelyonhydrophobicityoftheprotein
2.Arylafunctionalgroupderivedfromanaromaticringoftenaphenylgroup.Arylgroupsofferincreasing
specificity,asproteinscanalsointeractwiththefunctionalgroupthroughbasestackinginteractions.
The type and concentration of salt used for binding and elution must be determined empirically for each
protein.Additionally,therefoldingandfunctionoftheproteinmustbeensuredafteritselution.
Like other forms of column chromatography, an optimum HIC procedure can require a great deal of
troubleshooting. Adjusting buffer conditions and stationary phase are required for each protein to ensure
optimalseparation.
Sizeexclusionchromatography
(SEC)
Problem
Size
exclusion
chromatography
separates
proteins
by
their
hydrodynamic
radius,
a
property determined by both
the size and shape of the
Proteindoesnotbind
molecule. Unlike the other
chromatographic procedures
described previously, proteins
do not bind to a stationary
phase in SEC. Instead,
proteins are separated by the
Proteindoesnotelute
speed at which they navigate
through an inert stationary
phase(Figure9).
SEC is a valuable method
most often used in the final
stepsofapurificationscheme
[34],
unfolded
protein
molecules [35], and truncated
morereliablemethodofbuffer
exchangethandialysis,asitis
Runmoreequilibration
bufferthroughcolumnand
reloadprotein
Ionicstrengthofbinding
bufferistoolow
Increaseionicstrengthof
buffer
Proteinaggregatedatthe
ionicstrengthused
Decreaseionicstrengthof
bufferortryadifferentsalt
Ionicstrengthofelution
bufferistoohigh
Decreaseionicstrength
Proteinaggregatedon
column
Adjustbufferconditionsfor
moreproteinstability
Retentionistoohigh
Tryadifferentstationary
phasethatoffersless
retention
Flowrateiseithertoofast
ortooslow
Adjustflowrate
Columnwasnotwashed
sufficiently
Washwithalowerionic
strengthbufferClean
stationaryphaseaccording
tomanufacturer
Proteinaggregatedon
column
Adjustbufferconditionsfor
moreproteinstability
Poorretentiononcolumn
Tryadifferentstationary
phasethatoffersgreater
retention
Proteinisunfoldedor
aggregated
Adjustbufferconditionsfor
moreproteinstability
Proteindidnotreturntoits
Proteinlosesactivityduring
nativeconformation
procedure
exchanging
buffer
components. Often times,
SEC is used as a faster and
Solution
Columnwasnot
equilibrated
Lowresolution
duetoitsabilitytodifferentiate
betweendifferentspeciesofa
protein. Oligomeric species
species [36] can all be
separated from the native
protein, while simultaneously
Cause
Acofactorrequiredfor
activitywasremoved
duringpurification
Trybindingwithadifferent
saltoratalowerionic
strengthIncludeadditives
forproteinstability
Addcofactor
Table8.Troubleshootinghydrophobicinteractionchromatography.
SummarizedfromGE.
https://www.labome.com/method/ProteinPurification.html
12/16
26/09/2016
ProteinPurification
compatiblewithmoresolvents
and requires less buffer. A single solvent is used throughout the entire SEC procedure, and commercially
availableSECstationaryphasesarecompatiblewithmostcommonlyusedbuffercomponents.
The type of stationary phase used and the length of the column both play critical roles in influencing the
resolution of proteins separated by SEC. Several types of stationary phases are commercially available, and
thebestoptiondependsonboththemolecularweightoftheproteinsrequiringseparationandtheconditions
underwhichtheseparationwillbeperformed.
TypeofStationary
Phase
Matrix
Features
Offersquickbuffer
exchangeandgroup
separation
Workswellfor
molecularweight
determination
Sephadex
Crosslinkeddextranand
epichlorohydrin
Autoclavable
Matrixcanshrinkin
certainsolvents
Specifictypesof
sephadexare
availableforusewith
organicsolvents
Separatesmolecules
overalarge
Sephacryl
Crosslinkedallyldextran
andN,Nmethylene
bisacrylamide
molecularweight
range
Autoclavable
Workswithaqueous
andorganicsolvents
Highrecovery
Separatesmolecules
Sepharose
Crosslinkedagarose
overalarge
molecularweight
range
Highrecovery
Cannotbeautoclaved
https://www.labome.com/method/ProteinPurification.html
Superdex is often the best
choice for typical SEC
procedures,asitiscompatible
with most solvents and offers
thehighestresolutionofallthe
availablestationaryphases.
Aswithotherchromatographic
methods,
there
are
advantages
and
disadvantages to SEC, and
the decision to use this
method depends on both the
protein and its downstream
application.
AdvantagesofSEC:
1.Providesbufferexchange
anddesalting.
2.Enablestheseparationof
similarspecies(e.g.,
truncationsandoligomers)
thatmightnotbe
separatedwithother
purificationtechniques.
3.Iscompatiblewithmany
solvents.
4.Doesnotdependonany
specificproteinproperty
forretentionandelution.
DisadvantagesofSEC:
1.Performanceisvery
sensitivetocolumn
packing.
2.Canhavenonspecific
interactionsbetween
13/16
26/09/2016
ProteinPurification
Workswithaqueous
andorganicsolvents
Superose
Highlycrosslinked
agarose
Autoclavable
Hydrophobic
interactionsbetween
proteinsandmatrixis
possible
Compatiblewith
viscoussolvents
proteinandresin,which
decreasesresolution.
3.Offerslowresolutionfor
complexproteinmixtures.
4.Samplemustbeloadedat
asmallvolumefor
adequateresolution.This
canbeproblematicfor
proteinsthatprecipitateat
highconcentrations.
Optimizing SEC conditions to
Workswithaqueous
andorganicsolvents
Superdex
Crosslinkedagarose
anddextran
Autoclavable
Highresolution
Highrecovery
Table9.Typesofcommerciallyavailablestationaryphasesforsize
exclusionchromatographyandfeaturesofeach.SummarizedfromGE
andSimgaAldrich.
achieve maximum resolution
is often timeconsuming, but
certain factors can have a
hugeimpactonresolution.
Conditions that increase
proteinresolutionwithSEC:
1.Minimizethesample
volume.Thesmallerthe
volume,thelessdiffuse
theelutedfractionswillbe.
2.Addsalttothebuffer.A
smallamountofsaltwill
helppreventnonspecific
interactionsbetweenthe
proteinandstationary
phase.Thiswillallowall
proteinmoleculestorun
consistentlyoverthe
lengthofthecolumn.
3.Useamoderateflowrate.
Runningacolumntoo
quicklywillnotallowtime
forsmallmoleculesto
enterthematrixpores,
whileaflowratethatistoo
slowallowsmoretimefor
samplediffusion.
4.Ensurethatsampleand
solventviscositiesare
similar.Adjustsample
conditionsto
approximatelymatchthat
oftherunningbuffer.
5.Adjustthecolumnlength.
Acolumnthatistooshort
willnotallowadequate
proteinseparation.
Alternatively,acolumnthat
https://www.labome.com/method/ProteinPurification.html
14/16
26/09/2016
ProteinPurification
istoolongwillleadto
diffusionoftheprotein
sample.
6.Repackthecolumn.
Columnpackingcanhave
ahugeeffectonprotein
resolution.
Ifparticlesarenotwelldispersedorifairbubblesaretrappedinthecolumn,navigationthroughthestationary
phasewillnotoccurproperly.Additionally,ifthecolumnisallowedtorundry,itmustberepacked.Poorcolumn
packingisoftentoblameforunexplainedlowresolution.
BecauseSECdoesnotseparateproteinsbasedoninteractionswithafunctionalgroup,allproteinsareeluted
under the same conditions and, thus, resolution can only be obtained for proteins of very different
hydrodynamic radius. Therefore, SEC is not an appropriate chromatographic method for initial stages of a
purification scheme when there are many contaminating proteins. SEC does, however, offer a quick and
reliablemethodforremovingsaltsorsmallmoleculesfromasamplebetweenearlyorintermediatestagesof
thepurification.Inthefinalstagesofpurification,whenonlytracecontaminantsexist,SECisavaluablemethod
forproteinisolationandexchangeintostoragebuffer.
ProteinElutionandAnalysis
MethodsofProteinElution
Proteinsthatbindtoastationaryphaseareelutedwithsolventconditionsthatdisruptthebindinginteractions.
Theseelutionconditionsvarybothbytypeofchromatographyandpropertiesoftheproteinofinterest.There
are three general methods of protein elution: batchwise elution, stepwise elution, and linear gradient elution.
The best method to choose depends on the type of chromatography being performed and the required
resolution.
1.Batchwiseasingleelutionconditiondisplacesallboundproteinsinasinglestep.Thismechanismworks
bestforchromatographicproceduresbasedonaveryspecificinteraction(i.e.,affinitychromatography).
Batchwiseelutiondoesnotofferanyresolution,butitisidealforgettingridofcontaminantsveryquickly.It
requirespriorknowledgeofbufferconditionsrequiredfordisplacementoftheproteinofinterest.
2.Stepwisemultiplebatchwiseelutionsareperformedsequentially,withmorestringentconditionsineach
step.Instepwiseelution,thenumberoffractionscollectedisdependentuponthenumberofsequential
elutionconditions.Stepwiseelutionprovidesbetterresolutionthanbatchwiseelution,butpoorerresolution
thanlineargradientelution.
3.LinearGradientmultipleconsecutivefractionsarecollectedwhileelutionconditionsareadjustedina
linearfashion.Alineargradientoffersthehighestresolutionforionexchangechromatographyand
hydrophobicinteractionchromatography.Typically,alargenumberofconsecutivefractionsarecollected.
Size exclusion chromatography does not require any of these elution methods, as no interactions occur
between the protein and the stationary phase. Protein is loaded onto the column, and a large number of
fractionsarecollecteduntilallproteinsareeluted,withoutalteringbufferconditionsthroughouttheprocedure.
Ideally,theelutionbufferofacolumniscompatiblewiththesubsequentcolumn,eliminatingtheneedforbuffer
exchangeordialysisbetweenpurificationsteps.
AnalysisofFractionPurity
https://www.labome.com/method/ProteinPurification.html
15/16
26/09/2016
ProteinPurification
Aftereachchromatographicseparation,fractionsmustbeanalyzedtodeterminethefractionsthatcontainthe
proteinofinterestandtherelativepurityofeachofthosefractions.Thisanalysisisnecessaryaftereachstep
todecidewhichfractionsshouldbepooledforsubsequentuse.Todeterminepurity,anassayisrequiredthat
canmeasuretheamountofaspecificproteinrelativetotheamountoftotalprotein.Thefollowingassaysare
routinelyusedforpurityanalysis:
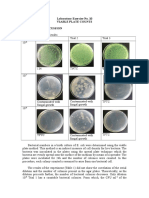
1. Sodiumdodecylsulfatepolyacrylamidegelelectrophoresis(SDSPAGE)adenaturinggelthatseparates
proteins based on size. Commercially available stains allow for a visual representation of all proteins in
the sample, offering a qualitative assessment of protein purity (Figure 10). SDSPAGE is not ideal for
highthroughputanalysisoffractionsandcantakeseveralhourshowever,itismostoftenusedbecause
itiseasy,inexpensive,andsuitableforanyprotein.
2.Spectroscopyamethodforanalyzingopticalpropertiesof
proteins.Thistechniqueforanalyzingproteinpurityisonly
suitedforproteins,suchascytochromeP450s,thathavea
uniquespectroscopicfeature.Proteinsinthisfamilyabsorb
lightatawavelengthwhereotherproteinsdonot(around420
nanometers[37]),soacomparisonofabsorbanceat420
nanometersversus280nanometers(thewavelengthatwhich
allproteinsabsorb),canprovideaquantitativemeasureof
proteinpurity.Thismethodisfastandhighthroughput,butonly
suitableforsomeproteins.
3.ProteinActivityanenzymatictestthatdependsontheprotein
ofinterest.Thismethodofassessingproteinpurityisoften
coupledwithanotherformofproteinconcentration
determinationtocalculateactivityrelativetototalprotein
concentration.Activityassaysareonlysuitableforproteins
withactivitythancaneasilybemonitoredinahighthroughput
Figure10.SDSPAGEofsamples
format,suchasproteases.Forsomeproteins,activityassays
collectingduringaproteinpurification provideafastandreliablemethodforproteindetection.Activity
scheme.Gelisstainedforthe
measurementisoftenidealforenzymes,becauseproteinthat
visualizationofallproteins.From
haslostactivitycanbeexcludedfromsubsequentuse.
http://www.omicsonline.org/.
PurifiedProteinStorage
Whenproteinsaredeemedpureenoughforuseinexperimentalstudies,theyshouldbestoredappropriately.
Theselectionofafinalstoragebufferisjustasimportantastheselectionofbuffersusedduringthepurification
scheme,andshoulddependonstabilityoftheproteinandconditionsrequiredforthedownstreamapplication
ofthepurifiedprotein.Oftentimes,sizeexclusionchromatographyisselectedasafinalstepinthepurification
scheme,asthestoragebuffercanbeusedinthischromatographicsteptoeffectivelyexchangethebuffer.The
purefractionscanbepooledforimmediatestorage.Alternatively,thefinalpooledfractionscanbedialyzedinto
theselectedbufferbeforestorage.
Proteinstorageconditionsdependontheproteinofinterest,andshouldbeoptimizedsotheproteinmaintains
structuralandfunctionalstabilityoverlongperiodsofstorage.Additivesareoftenincludedinthestoragebuffer
to enhance the lifetime of purified proteins under storage conditions, and trial and error is often required to
determineoptimumconditions,aseveryproteinbehavesdifferently.
https://www.labome.com/method/ProteinPurification.html
16/16
You might also like
- Gene Regulation of Prokaryotic TranscriptionDocument17 pagesGene Regulation of Prokaryotic TranscriptionAyu Tri AgustinNo ratings yet
- Instrumental Chemical Analysis ManualDocument39 pagesInstrumental Chemical Analysis ManualparthabakalNo ratings yet
- Dna RepairDocument20 pagesDna RepairEaron Van JaboliNo ratings yet
- Unit 3Document18 pagesUnit 3harshbharatdesai_783No ratings yet
- Mammalian Histology AssignmentDocument9 pagesMammalian Histology AssignmentSana Sultana100% (1)
- Isolation of Bacterial Plasmid DNA (Compatibility Mode)Document18 pagesIsolation of Bacterial Plasmid DNA (Compatibility Mode)Khandoker Faisal100% (1)
- Using Excel For Soil TestingDocument18 pagesUsing Excel For Soil TestingQaiser Abbas100% (3)
- lectut-BTN-303-pdf-Protoplast Technology PDFDocument12 pageslectut-BTN-303-pdf-Protoplast Technology PDFqwertNo ratings yet
- Administrative Theory in the 21st CenturyDocument132 pagesAdministrative Theory in the 21st CenturyRaja Mohan Gopalakrishnan83% (6)
- Enzyme Activity and AssaysDocument6 pagesEnzyme Activity and Assaysapi-318629889No ratings yet
- Cell and Tissue Culture TechniquesDocument44 pagesCell and Tissue Culture Techniquesdrashti shah100% (1)
- Module-3 (Theory) Upstream and Down Stream Component of A Fermentation Process PDFDocument6 pagesModule-3 (Theory) Upstream and Down Stream Component of A Fermentation Process PDFAnonymous AgGBWdNo ratings yet
- Gas ChromatographyDocument6 pagesGas ChromatographyPriyank ShahNo ratings yet
- Paradigms of Public AdministrationDocument10 pagesParadigms of Public AdministrationRaja Mohan Gopalakrishnan100% (1)
- Introduction To AC CircuitsDocument5 pagesIntroduction To AC CircuitsKeerthi Keerthana100% (1)
- Biology Tutorial (Set 1)Document18 pagesBiology Tutorial (Set 1)Jin MingNo ratings yet
- Basics of Cell Culture in 40 CharactersDocument33 pagesBasics of Cell Culture in 40 CharactersSai SridharNo ratings yet
- Cell Biology Lab Manual for B.Tech BiotechnologyDocument31 pagesCell Biology Lab Manual for B.Tech BiotechnologyVenus Divinagracia0% (2)
- The Theory of Public Administration (Presentation)Document191 pagesThe Theory of Public Administration (Presentation)Irfan WaheedNo ratings yet
- Lesson 1: How Does The Concept of Atomic Number Led To The Sysnthesis of New Elements in The LaboratoryDocument17 pagesLesson 1: How Does The Concept of Atomic Number Led To The Sysnthesis of New Elements in The LaboratoryAngelica Bag-ao100% (1)
- Proteomics IntroductionDocument39 pagesProteomics Introductionnariel67% (3)
- The Structure and Function of MacromoleculesDocument70 pagesThe Structure and Function of MacromoleculesLustried Nadyang100% (1)
- Activity of EnzymesDocument2 pagesActivity of EnzymessachithudaraNo ratings yet
- CELL PARTS and PROKARYOTESDocument6 pagesCELL PARTS and PROKARYOTESHarry ParconNo ratings yet
- Factors Affecting Enzyme ActivityDocument10 pagesFactors Affecting Enzyme ActivitygauravpriyaNo ratings yet
- Use of Micropippettor and SpectrophotometerDocument6 pagesUse of Micropippettor and SpectrophotometerMichelleNo ratings yet
- Enzyme Inhibitors Explained: Reversible vs IrreversibleDocument5 pagesEnzyme Inhibitors Explained: Reversible vs IrreversiblebelabelawNo ratings yet
- Prokaryoticneukaryoticgenome 160514180458 PDFDocument7 pagesProkaryoticneukaryoticgenome 160514180458 PDFrahul ssgiNo ratings yet
- Yeast Transgenic PlantsDocument5 pagesYeast Transgenic PlantsTooba Iqbal67% (6)
- Sutherland 1991Document7 pagesSutherland 1991Isal AbdussalamNo ratings yet
- RA 1425: Rizal LawDocument7 pagesRA 1425: Rizal LawRona LlamosoNo ratings yet
- Federis-3a-Act 1 Ra 1425Document5 pagesFederis-3a-Act 1 Ra 1425Faye Domini FederisNo ratings yet
- ABT 227 - Course Outline Introduction To Molecular Biology (3335)Document1 pageABT 227 - Course Outline Introduction To Molecular Biology (3335)justevansiNo ratings yet
- Factors Affecting Enzyme ActivityDocument32 pagesFactors Affecting Enzyme ActivityHus Arif100% (1)
- Marketing Concept, Customer Value, and ResearchDocument8 pagesMarketing Concept, Customer Value, and ResearchWilliams TaiwoNo ratings yet
- CELL CULTURE LINESDocument31 pagesCELL CULTURE LINESRamesh BeniwalNo ratings yet
- Biomolecules and Cells:: Mr. Derrick Banda MSC, BSCDocument69 pagesBiomolecules and Cells:: Mr. Derrick Banda MSC, BSCAmon Sangulube100% (1)
- CLSMDocument27 pagesCLSMprinceamitNo ratings yet
- 11th Biology Question BankDocument14 pages11th Biology Question BankJay senthilNo ratings yet
- Laboratory Exercise No. 10 Viable Plate Counts Results and DiscussionDocument3 pagesLaboratory Exercise No. 10 Viable Plate Counts Results and Discussionvanessa olga100% (1)
- Intracellular TransportDocument66 pagesIntracellular TransportalvitakhoridatulNo ratings yet
- Sampling and Isolation of Bacteria From SoilDocument9 pagesSampling and Isolation of Bacteria From SoilKishmala BatoolNo ratings yet
- Unit 3 The Three Dimensional Structure of ProteinsDocument20 pagesUnit 3 The Three Dimensional Structure of ProteinsPatricia OrtizNo ratings yet
- Lec04 MicroDocument13 pagesLec04 MicroMayurdhvajsinh JadejaNo ratings yet
- BCH 314 Tutorial 1 SolutionsDocument9 pagesBCH 314 Tutorial 1 SolutionsvictorNo ratings yet
- Animal Cell CultureDocument33 pagesAnimal Cell CultureMd. Babul AktarNo ratings yet
- NSCI 115: Chemical Principles of NANO I Lab 1: UV-Vis Spectroscopy and Beer-Lambert LawDocument6 pagesNSCI 115: Chemical Principles of NANO I Lab 1: UV-Vis Spectroscopy and Beer-Lambert LawIsaac SnitkoffNo ratings yet
- Cell Counting HemocytometerDocument4 pagesCell Counting HemocytometerJuan Dela CruzNo ratings yet
- MSC MPlil ZoologyDocument18 pagesMSC MPlil ZoologyYougesh KumarNo ratings yet
- Carbohydrates Classification and ReactionsDocument31 pagesCarbohydrates Classification and ReactionsAlviro CossemeNo ratings yet
- cDNA Libraries and Gene CloningDocument8 pagescDNA Libraries and Gene CloningRoberto RomeroNo ratings yet
- Biology Notes (Proteins)Document9 pagesBiology Notes (Proteins)Teo Jia Ming Nickolas100% (1)
- Rizal's Life and Works Study GuideDocument5 pagesRizal's Life and Works Study GuideGabrielle Beatrice TroncoNo ratings yet
- Carbohydrates: Definition, Structure, Properties and ClassificationDocument45 pagesCarbohydrates: Definition, Structure, Properties and ClassificationKiya AlemuNo ratings yet
- Immobilized Enzyme Kinetics Techniques and ApplicationsDocument92 pagesImmobilized Enzyme Kinetics Techniques and ApplicationsVNo ratings yet
- Separation Techniques For ProteinsDocument2 pagesSeparation Techniques For ProteinsRobert Dominic GonzalesNo ratings yet
- Fundamentals of BiochemistryDocument37 pagesFundamentals of BiochemistryTamoor TariqNo ratings yet
- Biology Laboratory Manual Series General Zoology Laboratory ManualDocument18 pagesBiology Laboratory Manual Series General Zoology Laboratory ManualAnnaNo ratings yet
- FicusDocument3 pagesFicusSugandha ShetyeNo ratings yet
- Principles of Cell Culture: A Brief History and OverviewDocument20 pagesPrinciples of Cell Culture: A Brief History and OverviewMicah Marie Cao AngeliasNo ratings yet
- Bacterial Optical Density MeasurementsDocument4 pagesBacterial Optical Density Measurementskrishnarao2010No ratings yet
- ELISA-Principle, Types and ApplicationsDocument4 pagesELISA-Principle, Types and ApplicationsSeema NegiNo ratings yet
- Importance of Tris EDTADocument15 pagesImportance of Tris EDTADarshana JuvekarNo ratings yet
- Bacteriology Lab 2002Document89 pagesBacteriology Lab 2002Romanda GreeneNo ratings yet
- 2.1. Cell Structure and Function of OrganellesDocument66 pages2.1. Cell Structure and Function of OrganellesNama Saya100% (1)
- A Theranostic and Precision Medicine Approach for Female-Specific CancersFrom EverandA Theranostic and Precision Medicine Approach for Female-Specific CancersRama Rao MallaNo ratings yet
- 40 BarrowDocument11 pages40 BarrowRaja Mohan GopalakrishnanNo ratings yet
- GlossaryDocument17 pagesGlossaryRaja Mohan GopalakrishnanNo ratings yet
- PCM June 2015 Fee ScheduleDocument2 pagesPCM June 2015 Fee ScheduleRikitaNo ratings yet
- 14 Chapter 3Document18 pages14 Chapter 3Raja Mohan GopalakrishnanNo ratings yet
- Chemistry Revision Pack - 2012Document15 pagesChemistry Revision Pack - 2012Raja Mohan GopalakrishnanNo ratings yet
- MTG Econ Lowres 2012Document196 pagesMTG Econ Lowres 2012Raja Mohan Gopalakrishnan100% (1)
- SPONSORING AGENCIES Financial Resources For Research & Development Sponsored AgenciesDocument118 pagesSPONSORING AGENCIES Financial Resources For Research & Development Sponsored AgenciesAshoka VanjareNo ratings yet
- Maths Formula SheetDocument2 pagesMaths Formula SheetRaja Mohan GopalakrishnanNo ratings yet
- Whither Constitutional BodiesDocument4 pagesWhither Constitutional BodiesRaja Mohan GopalakrishnanNo ratings yet
- Thesis Topic IrwinDocument15 pagesThesis Topic IrwinRaja Mohan GopalakrishnanNo ratings yet
- Bill Summary - Unlawful Activities - Prevention - Amendment Bill, 2011 v3Document1 pageBill Summary - Unlawful Activities - Prevention - Amendment Bill, 2011 v3Raja Mohan GopalakrishnanNo ratings yet
- Chemistry of Graphene and Its Technological Applications-Prof - LohDocument39 pagesChemistry of Graphene and Its Technological Applications-Prof - LohRaja Mohan GopalakrishnanNo ratings yet
- Types of RocksDocument19 pagesTypes of RocksAbdul Moeed KalsonNo ratings yet
- Reality of Divide and RuleDocument21 pagesReality of Divide and RuleRaja Mohan GopalakrishnanNo ratings yet
- 044 Kay ComplexDocument6 pages044 Kay ComplexRaja Mohan GopalakrishnanNo ratings yet
- Chemistry of Graphene OxideDocument13 pagesChemistry of Graphene Oxidenguyendktu100% (1)
- Burn 1977-A New Method For The Simultaneous Determination of The Size and Shape of Pore The ThermoporometryDocument40 pagesBurn 1977-A New Method For The Simultaneous Determination of The Size and Shape of Pore The ThermoporometryMin WuNo ratings yet
- Systems Approach TheoryDocument13 pagesSystems Approach TheoryRaja Mohan GopalakrishnanNo ratings yet
- Elements of The Organisation Through The Lens of System TheoryDocument16 pagesElements of The Organisation Through The Lens of System TheoryRaja Mohan GopalakrishnanNo ratings yet
- Systems - MilletDocument12 pagesSystems - MilletPèdjoeang SèdjatiNo ratings yet
- Evolution of Indian Administration & Key Principles of Kautilya's ArthashastraDocument1 pageEvolution of Indian Administration & Key Principles of Kautilya's ArthashastraRaja Mohan GopalakrishnanNo ratings yet
- Systems Theory and MCS-TNDocument9 pagesSystems Theory and MCS-TNonebyzerooutlookNo ratings yet
- Itish Involvementin IndiaDocument25 pagesItish Involvementin IndiaRaja Mohan GopalakrishnanNo ratings yet
- Modern India Revolt of 1857 Detailed Notes SampleDocument10 pagesModern India Revolt of 1857 Detailed Notes SampleSmriti SinghaniyaNo ratings yet
- Geography Extended Answer Notes SustainabilityDocument2 pagesGeography Extended Answer Notes SustainabilityRaja Mohan GopalakrishnanNo ratings yet
- Most Stable Nuclei Among Isobaric FamilyDocument1 pageMost Stable Nuclei Among Isobaric FamilyFast FeneNo ratings yet
- P525/2 Chemistry Paper 2: Uganda Advanced Certificate of Education Page 1Document8 pagesP525/2 Chemistry Paper 2: Uganda Advanced Certificate of Education Page 1ArthurNo ratings yet
- Photon and Quantized EnrgyDocument15 pagesPhoton and Quantized EnrgykayNo ratings yet
- Physics: AnswersDocument23 pagesPhysics: AnswersKrishna ChoureyNo ratings yet
- Q2 Chem 1 Molecular Geometry HandoutsDocument1 pageQ2 Chem 1 Molecular Geometry Handoutsmikomira21No ratings yet
- Central Angles PDFDocument8 pagesCentral Angles PDFJoseph LeeNo ratings yet
- Lab ReportDocument6 pagesLab Report'Izzad AfifNo ratings yet
- Technological Institute of the Philippines Mechanics of Deformable BodiesDocument19 pagesTechnological Institute of the Philippines Mechanics of Deformable BodiesChavin StormNo ratings yet
- Chem-Bonding AnswersDocument31 pagesChem-Bonding AnswersSahaj SinghNo ratings yet
- (Landolt-Börnstein - Group IV Physical Chemistry 5 J - Physical Chemistry) B. Predel (Auth.), O. Madelung (Eds.) - Pu-Re - Zn-Zr-Springer-Verlag Berlin Heidelberg (1998) PDFDocument584 pages(Landolt-Börnstein - Group IV Physical Chemistry 5 J - Physical Chemistry) B. Predel (Auth.), O. Madelung (Eds.) - Pu-Re - Zn-Zr-Springer-Verlag Berlin Heidelberg (1998) PDFNoe Rivera ONo ratings yet
- U2 - L10 Stress Isobar or Pressure Bulb PDFDocument6 pagesU2 - L10 Stress Isobar or Pressure Bulb PDFศิวาเวช อบมาNo ratings yet
- Dynamics of Rigid BodiesDocument14 pagesDynamics of Rigid BodiescharespiadalcordoNo ratings yet
- Course Code Course/Subject Name Credits CHC303 Fluid Flow 4Document3 pagesCourse Code Course/Subject Name Credits CHC303 Fluid Flow 4sddalviNo ratings yet
- Model Questions PhysicsDocument10 pagesModel Questions PhysicsCharis Israel AnchaNo ratings yet
- Measuring Load and Torque with Strain Gauge Load Cells and DynamometersDocument12 pagesMeasuring Load and Torque with Strain Gauge Load Cells and DynamometersAbigith BabyNo ratings yet
- CarbenesDocument25 pagesCarbenesmillinagi95No ratings yet
- Pentode Straped TriodeDocument32 pagesPentode Straped Triodebigpriap100% (1)
- 2-15-1466427029-1. Electrical - Ijeee - Coordinated Effect of Power System Stabilizer - Rampreet ManjhiDocument10 pages2-15-1466427029-1. Electrical - Ijeee - Coordinated Effect of Power System Stabilizer - Rampreet Manjhirobertovm2002No ratings yet
- Fundamental Parameters of Antenna (1) .PPSXDocument25 pagesFundamental Parameters of Antenna (1) .PPSXవేలుసామి లింగాసామిNo ratings yet
- 02 Statics of Rigid Bodies 01 ConceptsDocument3 pages02 Statics of Rigid Bodies 01 Conceptsxaaabbb_550464353No ratings yet
- Morison EquationDocument8 pagesMorison Equationmailnewaz96770% (1)
- Snakes and Ladders in Chemistry (Eamcet Special)Document11 pagesSnakes and Ladders in Chemistry (Eamcet Special)sriniwaas chhari thNo ratings yet
- Standard Heavy Duty Limit Switches FD/FP/FL: Options and Ordering CodesDocument9 pagesStandard Heavy Duty Limit Switches FD/FP/FL: Options and Ordering CodesNisar AhmedNo ratings yet
- Quest Review 4 Electric Force, Magnetic Fields KEYDocument16 pagesQuest Review 4 Electric Force, Magnetic Fields KEYJ PNo ratings yet
- Dom TutorialDocument7 pagesDom TutorialsrishashankNo ratings yet